 Recent Advances in the Genetics of Vocal Learning
Recent Advances in the Genetics of Vocal Learning
Michael C. Condro
Molecular, Cellular and Integrative Physiology Interdepartmental Program, University of California, Los Angeles
Stephanie A. White
Department of Integrative Biology and Physiology, University of California, Los Angeles
Reading Options:
Continue reading below, or:
Read/Download PDF | Add to Endnote
Abstract
Language is a complex communicative behavior unique to humans, and its genetic basis is poorly understood. Genes associated with human speech and language disorders provide some insights, originating with the FOXP2 transcription factor, a mutation in which is the source of an inherited form of developmental verbal dyspraxia. Subsequently, targets of FOXP2 regulation have been associated with speech and language disorders, along with other genes. Here, we review these recent findings that implicate genetic factors in human speech. Due to the exclusivity of language to humans, no single animal model is sufficient to study the complete behavioral effects of these genes. Fortunately, some animals possess subcomponents of language. One such subcomponent is vocal learning, which though rare in the animal kingdom, is shared with songbirds. We therefore discuss how songbird studies have contributed to the current understanding of genetic factors that impact human speech, and support the continued use of this animal model for such studies in the future.
Keywords: Autism, Basal ganglia, Cntnap2, FoxP1, FoxP2, KE family, Speech, Vocal learning, Zebra finch
Author Note: Author Note: Correspondence concerning this article should be addressed to Stephanie A. White, PhD, Department of Integrative Biology and Physiology, University of California, Los Angeles, 610 Charles E. Young Dr. East, Los Angeles, CA 90095-7239. E-mail: sawhite@ucla.edu.
Introduction
Vocal learning, which includes the ability to imitate sounds with one’s voice, is a rare trait in the animal kingdom. To date, only a few groups of mammals have demonstrated a capacity for vocal learning. These include certain species of echolocating bats, cetaceans, pinnipeds, elephants, and of course, humans (Fitch, 2012; Knornschild, Nagy, Metz, Mayer, & von Helversen, 2010; Stoeger et al., 2012). Outside of mammals, three groups of birds are capable of learning a portion of their vocalizations, namely hummingbirds, parrots, and songbirds, the last of which make up about half of all bird species (Reiner et al., 2004). The disparate pattern of vocal learning across taxa is characteristic of convergent evolution. A parsimonious explanation is thus that preadaptations for vocal learning emerged from non-learning ancestors of each taxon (Fitch, 2011). These preadaptations are likely genetically encoded, which suggests that despite the distant relationships between vocal learners, there are some common genetic factors. One 2013), even though these disorders are also characterized by language deficits. In contrast, within a sample of dyslexic children and their unaffected relatives, a single nucleotide polymorphism (T vs. C) in an intron of FOXP2, identified as rs7782412, was correlated with nonword repetition (NWR) score (Peter et al., 2011), with the major allele (T, frequency of 0.558) being associated with impairment on this task. Since dyslexia is associated with impairments of written, but not spoken, language (Lyon, Shaywitz, & Shaywitz, 2003), these data suggest that FOXP2 aberrations affect language processing as well as spoken motor ability. Notably, language processing deficits and low verbal IQ are symptomatic in the KE family as well (Vargha-Khadem, Watkins, Alcock, Fletcher, & Passingham, 1995), though it is unclear whether these traits are directly related to the FOXP2 mutation, or are sequelae of DVD.
FOXP2 Function in the Developing Brain
In all animals, the FOX family of transcription factors is involved in regulating biological processes that affect embryogenesis and tissue development, as well as processes underlying adult cancer and aging (Benayoun, Caburet, & Veitia, 2011; Carlsson & Mahlapuu, 2002). FoxP1, 2, and 4 are expressed in embryonic neural tissues (Lu, Li, Yang, & Morrisey, 2002; Shu et al., 2001), and may therefore mediate neurogenesis and/or differentiation. Experimental reduction of Foxp2 in the cortex of embryonic mice through either shRNA or overexpression of the dominant negative KE form of FoxP2 repressed the transition from radial precursor to immediate neuronal progenitor, resulting in decreased cortical neurogenesis (Tsui, Vessey, Tomita, Kaplan, & Miller, 2013). Interestingly, overexpression of human FOXP2 increases neurogenesis, whereas overexpression of murine Foxp2 does not. These data indicate that human FOXP2 exerts a greater neurogenic effect, which is perhaps significant for the construction of the brain, including neural circuits involved in language processing. Foxp2 (here indicating the mouse form of the protein by capitalizing only the first letter, whereas the human form contains all capitals, and camel case for all other chordates) (Kaestner, Knochel, & Martinez, 2000) in conjunction with Foxp4, appears to promote neurogenesis by regulation of N-cadherin (Rousso et al., 2012). In embryonic chick and mouse spinal cord, overexpression of either FoxP increases the release of neural progenitors from the neuroepithelium, whereas knockdown of both prevents this release. These effects have yet to be tested in the cortex.
Another mechanism whereby FoxP2 may promote the development of vocal learning circuitry is through neurite development, especially during embryogenesis. A recent gene ontology study using Foxp2-ChIP and expression arrays found that Foxp2 targets related to neurite development are enriched (Vernes et al., 2011). Using ex vivo neuronal cultures, this study found that expression of wild type Foxp2 accelerates neurite growth, whereas expression of the KE mutant form has the opposite effect. Ectopic expression of Foxp2, achieved by removing the 3’UTR, which includes its regulatory elements, delays neurite outgrowth in vitro, though by seven days neurites form properly (Clovis, Enard, Marinaro, Huttner, & De Pietri Tonelli, 2012).
FOXP2 regulates gene activity by binding to DNA either as a homodimer, or by heterodimerizing with FOXP1 or FOXP4. There are six known isoforms of FOXP2 (Figure 1), two of which are truncated and lack FOX domains (Bruce & Margolis, 2002). The truncated forms, referred to as FOXP2.10+ due to their alternate splicing at exon 10 (Figure 1), do not localize to the nucleus, but may still dimerize with other FOXP2 isoforms (Vernes et al., 2006). Therefore, it is hypothesized that FOXP2.10+ forms act as posttranslational regulators of FOXP2 activity. FOXP2 can also interact with C-terminal binding protein (CtBP) to repress transcription (Li et al., 2004). A new association has been identified between FOXP2 and the gene protection of telomeres 1 (POT1; Tanabe, Fujita, & Momoi, 2011). In cell culture, when POT1 is expressed alone or coexpressed with the KE dominant negative mutation (R553H) of FOXP2, it is not localized in the nucleus. Only when POT1 is coexpressed with wild type FOXP2 is nuclear localization observed. Loss of POT1 can elicit a DNA damage response and cause cell arrest (Hockemeyer, Sfeir, Shay, Wright, & de Lange, 2005). FOXP2, in conjunction with POT1, could therefore affect cell cycling during development. The human phenotype exhibited by the KE mutation may be partly mediated by the inability of the mutant FOXP2 to associate with POT1, thereby disrupting cell cycling during the development of neural tissues subsequently necessary for vocal learning (Tanabe et al., 2011).
Molecular Phylogeny of FoxP2
FoxP2 is highly conserved across species, particularly in the zinc finger and DNA-binding FOX domains (Figure 1). Two amino acid differences between humans and chimpanzees (303N and 325S in the human isoform; Figure 1) are unique to humans among living primates (Enard et al., 2002). Interestingly, these substitutions are shared with extinct hominids such as Neanderthals (Green et al., 2010; Krause et al., 2007; Reich et al., 2010), for whom the capability for language is still uncertain (Benítez-Burraco & Longa, 2012). Between the zebra finch and human isoforms, there are only five additional substitutions, including one in the zinc finger domain, which is conserved in primates and rodents, but differs in the zebra finch ortholog (Teramitsu, Kudo, London, Geschwind, & White, 2004). Importantly, the DNA binding region is conserved between zebra finches and humans, including the arginine residue corresponding to position 553 in humans that is the site of the KE mutation. There is a considerable amount of homology (>80%) in the zinc finger, leucine zipper, and DNA-binding domains between human FOXP2 and the single FoxP ortholog of fruit flies and honeybees, from which it is believed the vertebrate FoxP family expanded (Kiya, Itoh, & Kubo, 2008; Scharff & Petri, 2011). As in vertebrates, invertebrate FoxP is predicted to be involved in procedural learning and communication, consistent with its neural expression and suggesting that it is most distinct from mammalian FoxP3, which is not associated with neural cell types (Scharff & Petri, 2011). FoxP2 is not well-conserved among echolocating bats nor between bats and other mammals, however, which has been postulated to be the result of a selection pressure on FoxP2 in bats for the evolution of echolocation (Li, Wang, Rossiter, Jones, & Zhang, 2007).
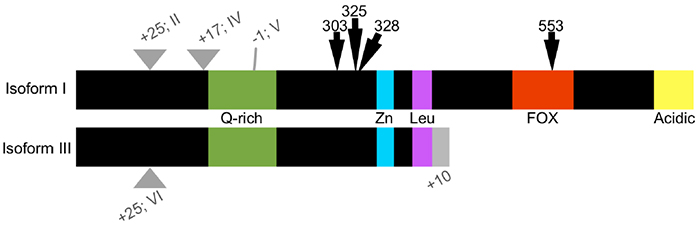
Figure 1. Schematic of human FOXP2 isoforms I–VI. FOXP2 is alternatively spliced as two major isoforms: the full-length isoform I and a truncated isoform III. Variations of either major isoform contain inserted or omitted amino acids (II, IV–VI), indicated here as the difference in number of amino acids (gray triangles). Both major isoforms possess a glutamine-rich (Q-rich) area, zinc finger (Zn) and leucine zipper (Leu) domains. Full-length isoforms of FOXP2 also possess a DNA-binding domain and an acid region on the C-terminus. Isoforms III and VI also have an additional 10 amino acids on the C-terminus that are not shared with the full-length isoforms. Arrows indicate amino acid substitutions between human and chimpanzee (303 and 325) or related to human speech disorders (328 and 553).
Songbird Studies of FoxP2
Humans are the only living animals that communicate with language (Berwick, Friederici, Chomsky, & Bolhuis, 2013), leaving no single animal model that sufficiently encapsulates every component of the behavior. However, facets of language are shared with other species. Vocal learning is one such facet that is shared with select groups of mammals, but as yet common laboratory models (e.g. rats, mice, nonhuman primates) fail to demonstrate this ability (Arriaga, Zhou, & Jarvis, 2012; Fitch, 2000; Mahrt, Perkel, Tong, Rubel, & Portfors, 2013). Rather, songbirds have been the principal animal models for vocal imitation in a laboratory setting (Panaitof, 2012). Vocal learning in both humans and songbirds relies on connections between the cortex, basal ganglia, and thalamus (Doupe & Kuhl, 1999). An advantage of the songbird model is that the neural structures responsible for vocal production and learning, called song production nuclei, are interconnected and anatomically distinct from the larger neurological subdivisions in which they reside, but are comprised of similar cell types. The song production nuclei are therefore assumed to function similarly to the circuits underlying other forms of procedural learning, but are dedicated to vocal learning. This feature of the songbird neuroanatomy has been incredibly useful for studies of vocal learning genes, many of which are discussed in this review. Among songbirds, zebra finches have been widely used due to their ease of breeding in captivity, as well as the sexual dimorphism of vocal learning (only males sing; Immelmann, 1969) and the song production system, which is incomplete in females (Konishi & Akutagawa, 1985; Nottebohm & Arnold, 1976).
FoxP2 mRNA expression is robust in the basal ganglia of humans and zebra finches (Teramitsu et al., 2004). In the zebra finch striatopallidal song nucleus, area X, FoxP2 transcript and protein levels correlate negatively with early morning singing. FoxP2 protein decreases in area X over the course of two hours when a male directs his songs at a female or when he practices them alone (Miller et al., 2008; Thompson et al., 2013); the latter is referred to as undirected singing. The transcript decreases during the course of two hours of undirected, but not directed, singing (Hilliard, Miller, Horvath, & White, 2012; Teramitsu & White, 2006; Teramitsu, Poopatanapong, Torrisi, & White, 2010). Down regulation of the mRNA is most potent in young birds engaged in sensorimotor learning (Teramitsu et al., 2010) when the more the juvenile practices, the lower his area X FoxP2 levels. This regulation appears largely due to motor activity, rather than auditory input, as levels also decrease in birds that have been deafened. However, there may be an additional auditory component to this phenomenon, as the degree of down regulation is only correlated with the amount of singing (Hilliard, Miller, Fraley, Horvath, & White, 2012) in juveniles that maintained their hearing (Teramitsu et al., 2010). The distinct behavioral regulation of the mRNA and protein suggests that there is post-transcriptional regulation of FoxP2, at least in the case of directed singing. In any case, both phenomena have been replicated at the two hour time point, namely that the protein levels decline with both directed and undirected singing, whereas the mRNA only declines with undirected song practice. Specifically, new findings show that microRNAs that target FoxP2 are up-regulated during undirected, but not directed, singing and lead to corresponding decreases in FoxP2 mRNA only for the former (Shi et al., 2013).
In their 2013 study of FoxP2 protein expression, Thompson et al. (2013) identified two categories of FoxP2-labeled neurons: those with large nuclei intensely labeled by the FoxP2 antibody, and those with smaller nuclei and weaker labeling. One possibility is that these subtypes represent different stages of maturation within a single population of medium spiny neurons (MSNs). Intensely labeled neurons may be younger neurons either in the process of migrating or already having migrated to area X, whereas weakly labeled neurons may be mature and integrated into the basal ganglia microcircuitry. The intensely labeled neurons peak in density within area X around 35 days and decline with age. The density of weakly labeled ‘mature’ neurons does not change with age. However, the density of these neurons in area X is behavioral context dependent. Adult males that sing for two hours in the morning exhibit a reduced density of weakly labeled neurons, a finding that replicates the behaviorally modulated levels of FoxP2 described by Fisher et al. (1998) and Miller et al. (2008).
In the zebra finch, experimentally induced reduction of FoxP2 at a developmental stage prior to the onset of vocal motor learning via injection of lentivirus containing an shRNA construct partially impairs the ability to learn the tutor’s song (Haesler et al., 2007). Though shRNA-injected young zebra finches are capable of producing sounds similar to those of their tutors, they consistently fail to accurately imitate the tutor’s song, often omitting or repeating individual syllables. Additionally, they are unable to accurately imitate the spectral characteristics and timing of the tutor’s song. During this period of song learning, new neurons expressing FoxP2, which are hypothesized to affect behavioral plasticity, migrate into area X (Rochefort, He, Scotto-Lomassese, & Scharff, 2007). Surprisingly, though, knockdown of FoxP2 does not prevent the proliferation of new neurons from the ventricular zone. It does, however, reduce the number of dendritic spines on MSNs, suggesting that FoxP2 affects neuronal plasticity without affecting proliferation and migration of new neurons (Schulz, Haesler, Scharff, & Rochefort, 2010). These data provide support for a functional role of FoxP2 in vocal learning subserved by basal ganglia circuits, in addition to mediating the development of the brain regions involved.
Mouse Models of Foxp2
Several mutant mice strains have been generated to study the effects of Foxp2 on brain morphology as well as vocal and nonvocal behaviors. In one such model, the two amino acids characteristic to humans (Enard et al., 2002) were changed to conform to the human sequence (Enard et al., 2009). The resulting mice have altered cortico-basal ganglia circuitry in the form of increased dendrite length in Foxp2-expressing bipolar spiny neurons in layer 6 of the primary motor cortex, MSNs in the striatum, and neurons in the parafascicular nucleus of the thalamus. Longterm depression (LTD) is increased in MSNs of the striatum, and dopamine concentrations are reduced in several brain regions, including the striatum (Reimers-Kipping, Hevers, Pääbo, & Enard, 2011). Despite also expressing the humanlike Foxp2 protein, dendrite lengths of amygdalar and cerebellar Purkinje neurons are unchanged. Purkinje cell LTD is also similar to control levels, which suggests that the humanlike Foxp2 impacts mainly basal ganglia microcircuits (Enard et al., 2009; Reimers-Kipping et al., 2011). In terms of behavior, the mutant mice exhibit decreased exploration, spend more time in groups, and as neonates emit ultrasonic vocalizations with reduced pitch and increased frequency modulation compared to control mice. Interestingly, FOXP2 knockout heterozygotes with a functional wild type allele have the opposite effects on dopamine levels and behavior (Enard et al., 2009).
Several mouse models have been generated to mimic FOXP2 mutations associated with human disorders. These knock-in mice include murine versions of the KE mutation (R552H; Fujita et al., 2008; Groszer et al., 2008), a similar mutation that results in an amino acid substitution at a different site within the DNA binding domain (N549K; Groszer et al., 2008), and a truncation (S321X) that fails to produce a protein, similar to a human mutation associated with speech impairment (Groszer et al., 2008). These loss of function knock-in mutations are lethal in homozygotes, with mice usually dying within the first month of life, though N549K homozygotes can survive for several months. All knock-in mutants have decreased cerebellar volume and Purkinje cell dendritic arbor (Fujita et al., 2008; Groszer et al., 2008), but otherwise no gross anatomical disturbances were observed in the rest of the brain. Homozygous knockout, R552H, and S321X mutant mouse pups make fewer ultrasonic distress calls, though there are mixed reports about the quality of these vocalizations (Fujita et al., 2008; Gaub, Groszer, Fisher, & Ehret, 2010; Groszer et al., 2008; Shu et al., 2005). Recently, Bowers, Perez-Pouchoulen, Edwards, & McCarthy (2013) investigated these calls using wild type rats and found qualitative and quantitative sex differences. Similar to mice, isolation calls are emitted from rat pups separated from their dam and trigger her to retrieve the pup back to the nest. The authors found that male pups call more frequently, at a lower pitch, and more quietly than do female pups. In turn, the dam responds differently to calls made by each sex, preferentially retrieving male before female pups. Male rat pups have more Foxp2 protein than female pups in several brain areas. Experimental reduction of Foxp2 by injection of siRNA into the ventricles during the first two days of life reverses this sex effect in calling behavior. Treated male pups call less frequently and at a higher pitch than control males. Notably, treatment of female pups with siRNA causes their vocalizations to become male-like, with higher frequency of calling, lower pitch, and lower amplitude. The authors posit that the reversal caused by Foxp2 siRNA is the result of a decrease in Foxp2 in males and a rebound-effect increase in females, although no evidence is provided for the latter. Interestingly, the dam retrieves siRNA-injected females before siRNA-treated males, providing evidence that the retrieval response of the dam depends on the vocal behavior rather than other sexually dimorphic characteristics. This study also finds that, in postmortem human brain tissue, there is more FOXP2 in the cortices of 4-year-old girls than age-matched boys, which coincides with gender-based language differences in children at this age. The authors posit that sex differences in brain FoxP2 levels correlate with the more ‘communicative’ sex in human and rodent species.
Since separation calls are not learned (Arriaga et al., 2012) and therefore are not analogous to human speech, studies in Foxp2 mutants examined other classical learned behavioral skills. One such skill was measured by Morris water maze place learning, in which mice were given four consecutive training trials each day for four days, after which the platform was moved and training began again (Santucci, 1995). Heterozygote knockout mice perform as well as wild types (Shu et al., 2005), indicating that this hippocampal-based learning task is not affected by loss of Foxp2. However, R552H mutants are impaired on the accelerating rotarod, a procedural learning task in which mice are placed on a rod that rotates around its axis at an increasing rate and the amount of time before the animal falls from the rod is recorded. Performance on the rotarod relies on basal ganglia activity, suggesting that R552H mutant mice have deficits in activity in these brain regions (French et al., 2012; Groszer et al., 2008). R552H heterozygous mutant mice have corresponding neurophysiological abnormalities, including reduced striatal LTD and increased cerebellar paired pulse facilitation (Groszer et al., 2008). In vivo electrophysiological recordings of these mice during the accelerating rotarod learning task show that striatal firing rate activity decreases in R552H mutants, whereas it increases in wild type, and temporal coordination is altered (French et al., 2012). Interestingly, these mutant mice can perform other striatal-based learning tasks, such as pressing a lever for a reward, equally well as controls. These data suggest that Foxp2 activity in the basal ganglia is involved in procedural learning tasks in nonvocal learning species, perhaps in a similar manner to vocal learning in humans and songbirds.
2. FOXP1
FoxP1 is the most similar molecule to FoxP2 and, perhaps not surprisingly, is also linked to human speech. As previously mentioned, FoxP1 and FoxP2 may form heterodimers that regulate transcription in areas where their expression overlaps (Li et al., 2004; Shu et al., 2001; B. Wang, Lin, Li, & Tucker, 2003). Initial support for a role of FOXP1 in vocal learning stems from a study of comparative gene expression in two vocal learners: humans and zebra finches. Unlike FoxP2, for which differential expression in song nuclei depends on behavior, FoxP1 signals constitutively ‘mark’ the song system, with mRNA enrichment in area X (in males), HVC, and RA relative to their surrounding tissues (Teramitsu et al., 2004). In humans, FOXP1 and FOXP2 are found in separate cortical layers: the former is found primarily in layers 2/3 with less expression in deeper layers, whereas the latter is primarily in layer 6 (Ferland, Cherry, Preware, Morrisey, & Walsh, 2003; Teramitsu et al., 2004). Both transcripts are expressed in the human striatum, similar to the expression pattern in the basal ganglia nucleus area X of songbirds. The possible co-regulation of transcription by FoxP members in the songbird song production system, and the comparative gene expression in humans suggested that FOXP1 also plays a role in human language (Teramitsu et al., 2004). Subsequently, Pariani, Spencer, Graham, & Rimoin (2009) reported the first human case of FOXP1 alteration and speech impairment, in which the patient had a large deletion in chromosome 3 including the FOXP1 gene. Speech delay was one of several deficits, which also included anatomical and neurological abnormalities. Shortly after this report, several similar cases were published in which patients with FOXP1 deletions presented cognitive deficits, motor control deficits, speech delay, and autism (Carr et al., 2010; Hamdan et al., 2010; Horn, 2012; Horn et al., 2010; O’Roak et al., 2011; Palumbo et al., 2013; Talkowski et al., 2012). In all the reported cases, however, the language impairment described was more consistent with speech delay than DVD. A screen of patients with DVD failed to identify FOXP1 as a risk factor (Vernes, MacDermot, Monaco, & Fisher, 2009). Though many of the phenotypes associated with mutations in FOXP1 and FOXP2 are non-overlapping, language impairment is common to both (Bacon & Rappold, 2012).
3. CNTNAP2
CNTNAP2 in Human Disease
Similar to the discovery of the relationship between FOXP2 and language through the KE family, a rare mutation in the contactin associated protein-like 2 (CNTNAP2) gene was discovered in a genetically related population of Old Order Amish children (Strauss et al., 2006). Some members of this group are afflicted with cortical dysplasiafocal epilepsy (CDFE). The disorder is characterized by the onset of seizures at about 2 years of age, mental retardation, hyperactivity, pervasive developmental delay or autism in the majority of cases, and language regression by the age of 3 in all cases. Patients with CDFE are homozygous for a deletion of a single base pair in CNTNAP2 exon 22, 3709delG. Subsequent to the initial association between CNTNAP2 mutation and CDFE, it was revealed that it is transcriptionally regulated by FOXP2. In chromatin immunoprecipitation (ChIP) assays, fragments of intron 1 of CNTNAP2 were bound by FOXP2 at the canonical binding sequence CAAATT (Vernes et al., 2008; Vernes et al., 2011). Mutation of these sites to CGGGTT prevented FOXP2 binding. Overexpression of FOXP2 in the human-derived neuroblastoma cell line SY5Y decreased CNTNAP2 transcription. To further investigate the relationship between CNTNAP2 and language ability, variants of the gene were screened in a cohort of families with SLI-afflicted members. Nine intronic SNPs between exons 13 and 15 of CNTNAP2 correlated with NWR scores. The one SNP most correlated, rs17236239, was also associated with expressive language score. Quantitative transmission disequilibrium testing (QTDT) confirmed a relationship between measures of language ability and four of these SNPs, but failed to confirm a relationship for rs7794745 in a new sample of families containing members with SLI (Newbury et al., 2011). None of the SNPs associated with language-related QTDT measures in a sample of families with dyslexia, indicating that there are separate factors that affect language ability.
Other common CNTNAP2 polymorphisms have been identified that associate with diagnoses of autism (Arking et al., 2008; Bakkaloglu et al., 2008), for which language impairment is a core deficit, and a language-related measure, age at first word (Alarcón et al., 2008). Interestingly, inherited CNTNAP2 polymorphisms that are associated with disease occur mainly in introns (Alarcón et al., 2008; Arking et al., 2008), suggesting either these SNPs are in linkage disequilibrium with yet unidentified markers in exons, or the SNPs themselves affect transcriptional regulation of the gene. Quantitative transmission disequilibrium testing revealed a association between the SNP rs2710102 and NWR (Peter et al., 2011). Thirteen de novo mutations in CNTNAP2 have been described in ASD patients that result in an amino acid change in the protein, eight of which were predicted to hinder function (Bakkaloglu et al., 2008). The de novo mutations, along with the CDFE mutation identified by Strauss et al. (2006), were investigated further to determine whether they did in fact affect protein function. HEK cells and rat hippocampal neurons were transfected with either wild type human CNTNAP2 or the mutant forms (Falivelli et al., 2012). The mutation associated with CDFE, 3709delG, causes a frameshift that results in the loss of the single transmembrane and intracellular domains of the protein (Figure 2, Strauss et al., 2006). This causes the normally membrane-bound protein to be secreted instead (Falivelli et al., 2012), presumably eliminating its normal functionality, and possibly introducing novel effects.. Another mutant, D1129H (Figure 2), also prevents surface expression of CNTNAP2, and instead the protein remains restricted to the endoplasmic reticulum, unable to move to the plasma membrane, interferes with the LNS4 domain of CNTNAP2, and is presumed to cause misfolding of the protein. Most other mutations investigated did not show restricted localization to the ER, though a mutation in a highly conserved amino acid, I869T (Figure 2), had less surface staining than the wild type form of the protein. Theoretically, mutations that interfere with intracellular trafficking of CNTNAP2 would also interfere with protein function. However, with the exception of 3709delG, these mutations do not always result in an autistic phenotype, indicating that other genetic, environmental, and developmental factors are involved in the presentation of the disorder.
CNTNAP2 Function in the Brain
Investigation of genes related to the formation of language-related brain areas revealed CNTNAP2 enrichment in the cortical superior temporal gyrus, associated with language processing and production (Abrahams & Geschwind, 2008). Moreover, CNTNAP2 is enriched in embryonic human frontal cortex, but not in rat or mouse at comparable stages of development. Not only do these data suggest a potential role for CNTNAP2 in the development of neural circuitry underlying language, they conform to the idea that this enrichment is relevant to vocal learning in humans, a behavior not shared with rodents.

Figure 2. Schematic of human CNTNAP2. CNTNAP2 consists of a single discoidin domain (DISC), four laminin-G domains (LamG), EGF repeats, a single transmembrane region (TM), and a putative protein 4.1 binding region (4.1m). CDFE indicates the subregion of the protein that is deleted in cases of cortical dysplasia-focal epilepsy in an Old Order Amish population (Li et al., 2004; Strauss et al., 2006). Arrows indicate two other amino acid changes associated with language impairment (869 and 1129).
The brains of healthy and autistic individuals homozygous for risk alleles rs7794745 and rs2710102 exhibit functional differences. Subjects with one or both risk variants exhibit increased activation of the frontal operculum and medial frontal gyrus relative to subjects homozygous for the non-risk allele (Whalley et al., 2011). Event-related brain potentials are altered during a language perception task in individuals carrying the rs7794745 risk allele (Kos et al., 2012). Scott-Van Zeeland, Abrahams, et al. (2010) investigated the correlation of risk allele rs2710102 with connectivity both within the medial prefrontal cortex (mPFC) and between other areas. In this study, subjects participated in a reward-based learning task in which they were presented with abstract images and were asked to assign them into either “Group 1” or “Group 2.” Upon correct classification, subjects were either given a monetary or social reward, or a “neutral” reward in which they were simply told whether or not they were correct. This experimental paradigm activates frontostriatal circuits (Scott-Van Zeeland, Dapretto, Ghahremani, Poldrack, & Bookheimer, 2010). Subjects with the CNTNAP2 risk allele rs2710102 exhibited increased local connectivity in the mPFC relative to subjects without the risk variant. This occurred in a genetically dominant fashion regardless of the autism phenotype of the risk allele carriers. In addition, risk allele carriers had less focused long-range connectivity between the mPFC and several other brain areas, as well as decreased lateralization, a result which is associated with autism-like behaviors. These data suggest that CNTNAP2 variants increase the risk of autism through alteration of frontal lobar connectivity.
Animal Models for Cntnap2
As yet, the most well-characterized function of Cntnap2 is to cluster voltage-gated potassium channels at juxtaparanodes of axons in the peripheral nervous system (Poliak et al., 2003). Recently, another potential function was discovered through an RNA interference (RNAi) survey of autism susceptibility genes (Anderson et al., 2012). Of the 13 genes included in the RNAi screen, Cntnap2 knockdown had the most pronounced effects on network activity in mouse hippocampal cultures. In mouse cortical cultures transfected with short hairpin RNA (shRNA) targeting endogenous Cntnap2, calcium transients from evoked synaptic responses were reduced in amplitude to approximately 70% of controls, though action potential frequency was not affected. Conversely, knockdown of the binding partner of Cntnap2, contactin 2, had the opposite effect, increasing the amplitude of the action potential. Cntnap2 expression level has no effect on neuronal excitability. Instead, the underlying cause of the action potential attenuation is a global decrease in synaptic transmission. Both excitatory and inhibitory evoked currents are reduced by the shRNA, as well as the frequency of miniature postsynaptic potentials, suggesting that the number of synaptic sites on affected neurons is reduced. This is further confirmed by changes to cellular morphology of transfected neurons. Cntnap2 knockdown results in shorter neurites with fewer branches, and dendritic spines with smaller spine heads. These data are evidence that Cntnap2 may affect the development of neurons by increasing the number of active synaptic sites and facilitating network activity.
Given the evidence for a role of CNTNAP2 in human speech, it may also function in birdsong (Panaitof, Abrahams, Dong, Geschwind, & White, 2010). In adult male zebra finches, Cntnap2 transcript is enriched in the robust nucleus of the arcopallium (RA) and the lateral magnocellular nucleus of the anterior nidopallium (LMAN), cortical nuclei in the song production system. Projection neurons from RA are similar to layer 5 pyramidal neurons in mammalian cortex whose axons descend below the telencephalon to synapse onto motor neurons (Jarvis, 2004), and LMAN shares similarities with the mammalian prefrontal cortex (Kojima, Kao, & Doupe, 2013). No such enrichment of Cntnap2 is observed in HVC (acronym used as a proper name), another song nucleus analogous to mammalian cortical layer 2/3 (Jarvis, 2004), and there is reduced expression in area X relative to the striatopallidum. Each song nucleus is comprised of similar cell types as those in the surrounding tissues, which suggests that the differential expression of genes within the song nucleus indicates a specific role for those genes in vocal learning and/or production. In contrast to males, adult females have moderate transcript levels in RA and LMAN. Female zebra finches have an underdeveloped area X that is not visible by common staining procedures (Balmer, Carels, Frisch, & Nick, 2009), but still Cntnap2 is uniform across the entire striatopallidum. Interestingly, in young females (<50d) Cntnap2 is enriched in RA to the same degree as for males, and declines to the level of the surrounding arcopallium with age. The reduction in gene expression coincides with the sensorimotor period of song learning in males, a time at which the male begins to practice singing. The percentage of cells expressing the protein in female RA decreases at this time point (Condro & White, 2014). This sexually dimorphic expression supports the hypothesis that Cntnap2 expression in RA is important for proper production of learned vocalizations in songbirds. According to this hypothesis, interference of Cntnap2 translation in male RA should disrupt song learning and/or production (Haesler et al., 2007).
As with Foxp2, mouse models of Cntnap2 risk variants may not capture language deficits associated with their respective disorders. However, they can be used to study other aspects of behavior and physiology that may impact future studies focused on vocal learning. Initially, outbred Cntnap2(-/-) mice were reported to have no gross anatomical or neurological abnormalities (Poliak et al., 2003). However, when these mice were crossbred with the C57BL/6J strain, subsequent generations exhibited neurological abnormalities similar to human patients with CDFE (Strauss et al., 2006), including epileptic seizures induced by mild handling starting before 6 months of age (Penagarikano et al., 2011). These knockout mice present neuronal migration abnormalities, with an increase in the incidence of ectopic neurons, a reduced number of inhibitory interneurons in the cortex and the striatum, along with impaired network synchrony in the cortex. Additionally, there is increased spontaneous inhibitory activity in cortical layers 2/3, disrupting the balance between inhibition and excitation (Lazaro, Penagarikano, Dong, Geschwind, & Golshani, 2012). These mice exhibit behavior similar to the human autistic phenotype, including repetitive motions, such as self-grooming and digging, behavioral inflexibility on learned tasks, such as the Morris water maze or T maze, decreased social activity with other mice, reduced nest building, and a decrease in the number of ultrasonic separation calls. Less frequent vocalizations could be symptomatic of impaired communication similar to language regression in autism, or alternatively due to a decreased motivation for maternal interactions, similar to social impairment in autistic children. The two hypotheses are not mutually exclusive, though the former is less likely, since this particular call type in mice is innate (Arriaga et al., 2012) and therefore not subject to regression. Interestingly, many of the behavioral deficits in the knockout mice can be partially rescued by treatment with risperidone, a medication used to treat the symptoms of autism (Penagarikano et al., 2011). However, the drug does not improve social interactions for the knockout mice. The effects of risperidone on communicative behavior have not yet been reported. Rescue by the drug of some of the effects of knocking out Cntnap2 further validates the relationship between Cntnap2 and autism. These knockout mice can be used to test other drugs to treat some of the symptoms of autism, though perhaps not language impairment. This model is especially pertinent to CDFE, for which the mutation renders CNTNAP2 nonfunctional. The more common polymorphisms associated with ASD and SLI risk lie in introns, creating a challenge to develop mouse models. A songbird model may offer an advantage in understanding the role of CNTNAP2 in language in that knockdown of Cntnap2 can be targeted to song nuclei, isolating its effects on vocal behavior.
4. Hepatocyte Growth Factor Signaling Pathway Genes
In keeping with the theme of FoxP2 as a molecular entry point into gene networks involved in speech and language, another class of FoxP2 target genes is implicated in language deficits. Three genes in the hepatocyte growth factor (HGF) signaling pathway are each targets of FOXP2 regulation and associated with disorders of human speech and language. The first is the HGF receptor tyrosine kinase MET (Bottaro et al., 1991), which has been linked to ASD (Mukamel et al., 2011). The second, also linked to ASD, is the urokinase plasminogen activator receptor (uPAR, or PLAUR when referring to the human gene; Campbell et al., 2007), which was long thought to indirectly activate HGF through its binding partner urokinase (Mars, Zarnegar, & Michalopoulos, 1993), though more recently this function has been challenged (Eagleson, Campbell, Thompson, Bergman, & Levitt, 2011; Owen et al., 2010). The third is sushi-repeat protein, X-linked 2 (SRPX2), a uPAR ligand (Royer-Zemmour et al., 2008) that also binds HGF (Tanaka et al., 2012), and may account for the HGF-mediated effects of uPAR signaling. SRPX2 is linked to language through association with childhood seizures of the Rolandic fissure, which can cause language disabilities (Roll et al., 2006). FOXP2 binds the promoter regions of all three genes and represses transcription (G. Konopka et al., 2012; Mukamel et al., 2011; Roll et al., 2010). Recent evidence suggests that FOXP2 regulation of SRPX2 affects synaptogenesis and vocalizations in mice (Sia, Clem, & Huganir, 2013). Similar to CNTNAP2, the distribution of MET in human fetal brain is complementary to that of FOXP2. In cultures of normal human neural progenitors and established cell lines, endogenous FOXP2 expression increases with maturity as MET decreases (G. Konopka et al., 2012). Notably, the KE mutant (R553H) fails to repress uPAR or SRPX2 (Roll et al., 2010). These data suggest that HGF signaling is altered in cases of language disorders associated with FOXP2. To date HGF itself has not been directly associated with a disorder relating to speech; however, given the association of these other HGF signaling pathway genes with language disorders, it would not be surprising if such an association were discovered.
MET was initially investigated as an autism susceptibility gene due to the similarity of neuroanatomical abnormalities attributed to loss of MET signaling in the cortex and those found in cases of autism (Campbell et al., 2006). A SNP in the promoter region of MET, rs1858830, was identified as a site associated with elevated risk of diagnosis of autism. The “C” variant at this site causes a reduction in transcription of the gene, and alters transcription factor binding relative to the non-risk “G” variant. The “C” variant is overrepresented in cases of ASD, associated with reduced MET protein in the cortex (Campbell et al., 2007; Campbell, Li, Sutcliffe, Persico, & Levitt, 2008) and social and communication impairments in cases of ASD (Campbell, Warren, Sutcliffe, Lee, & Levitt, 2010). In healthy human embryonic brains, MET is enriched in the temporal cortex, an area involved in language processing, and to a lesser degree in the hippocampus and occipital cortex (Mukamel et al., 2011). HGF signaling through MET promotes development of cortical projection neurons (Eagleson et al., 2011). In microarray analysis, MET has been identified as a member of a gene module correlated with differentiation, particularly with axon guidance (G. Konopka et al., 2012). Though protein levels are dynamic during development, a peak of expression coincides with increased development of neurites and synapse formation, suggesting a role for MET in neuronal connectivity (Judson, Bergman, Campbell, Eagleson, & Levitt, 2009). MET is expressed in axon tracts of projection neurons of the neocortex, including those that descend into the striatum, consistent with the hypothesis that MET is a factor in development of neural circuits, which when perturbed, leads to symptoms of ASD and language impairment.
In a screen for other ASD-related genes in the MET signaling pathway, a SNP in the promoter region of PLAUR, rs344781, was identified as a risk factor for autism diagnosis with an interaction effect with MET rs1858830. uPAR knockout mice have been generated, but thus far studies have focused on the effects of knockout on neural migration and seizure activity. Whereas MET seems to promote cortical projection neuron migration and growth, uPAR seems to affect inhibitory neurons in much the same manner, though the mechanism remains unclear (Eagleson et al., 2011). Homozygous knockouts exhibit spontaneous seizures as well as a reduction of parvalbumin-positive interneurons in the anterior cingulate and parietal cortices (Eagleson, Bonnin, & Levitt, 2005; Powell et al., 2003). The loss of inhibitory interneurons may affect the balance of excitation and inhibition, a phenomenon associated with autism (Eagleson et al., 2011). Interestingly, uPAR may be absent in birds (NCBI search, BLAST), suggesting that it is not common to all vocal learning species. There may be a different molecule in songbirds that replaces uPAR function. Though uPAR was originally thought to be involved in the activation of HGF required for binding to MET (Mars et al., 1993), recent evidence suggests that uPAR and its binding partner urokinase contribute very little to the process, and rather other serine proteases are responsible for HGF activation (Owen et al., 2010). Phenotypic differences in uPAR and MET knockout mice support this hypothesis (Eagleson et al., 2011). However, uPAR is involved in several signaling cascades independent of MET (Blasi & Carmeliet, 2002), any of which may be related to autism or language impairment.
SRPX2 is a chondroitin sulfate proteoglycan that binds to both HGF and uPAR (Royer-Zemmour et al., 2008; Tanaka et al., 2012). Mutations in SRPX2 can result in seizures originating in the Rolandic fissure, which can lead to abnormal brain morphology in the form of polymicrogyria, and are associated with oral and speech dyspraxia and cognitive impairment (Roll et al., 2006). One such mutation, resulting in a tyrosine-to-serine substitution at position 72, is related to both Rolandic seizures and orofacial and fine motor impairment. The substitution occurs in a region thought to affect protein–protein interactions. In this same region, a site at position 75 is highly conserved among primates, but has changed in humans since the split from chimpanzees, suggesting an evolutionary mechanism for human speech (Royer et al., 2007), reminiscent of the amino acid substitutions in FOXP2 between the two species (Enard et al., 2002). As mentioned previously, new evidence has emerged for the role of SPRX2 in mouse vocalizations (Sia, Clem, & Huganir, 2013). Other chondroitin sulfate proteoglycans are involved in formation of perineuronal nets, which can affect plasticity of sensory systems (McRae, Rocco, Kelly, Brumberg, & Matthews, 2007). In songbirds, development of perineuronal nets around song nuclei correlates with the development of song, and it is hypothesized that destruction of these nets permits the reopening of critical period for song learning after crystallization (Balmer et al., 2009). It is possible, therefore, that SRPX2 is involved in similar processes, which could affect learned vocalizations in humans and songbirds alike.
5. Stuttering Genes
Stuttering, or stammering, is a condition in which speech is interrupted by involuntary repetitions of syllables or words, prolongation of syllables, or pauses during speech. Inheritance patterns strongly suggest a multifactorial genetic basis for the disorder, with relatively little environmental influence (Kang et al., 2011; Kraft & Yairi, 2012). However, it was not until recently that any specific gene was identified as a factor in stuttering. Genome-wide linkage revealed a locus of disequilibrium on chromosome 12 for stuttering (Riaz et al., 2005), which was investigated more closely in a large pedigree, identified only as Family PKST72, in which roughly half of the living members stutter (Kang et al., 2010). Genotyping in this pedigree revealed a relationship with a SNP, (G3598A), which causes a glutamine-to-lysine amino acid substitution in a gene encoding a subunit of N-acetylglucosamine-1-phosphate transferase (GNPTAB). The ‘A’ variant was more common in stuttering family members, and family members homozygous for the ‘G’ variant were much less likely to stutter. Unlike FOXP2 in the KE family, though, G3598A exhibits some phenotypic plasticity, in that not every family member with an ‘A’ variant stutters, and some family members homozygous for the ‘G’ variant do stutter. Sex has been previously shown to be a factor in recovery of stuttering, with females being four times more likely to recover (Ambrose, Cox, & Yairi, 1997). Such may be the case for the two female non-stuttering family members homozygous for the ‘A’ variant (Kang et al., 2010). Three more amino acid changes in GNPTAB were associated with stuttering in a broader population sample, as well as three others found in GNPTG, another subunit of the phosphotransferase, and three more mutations in N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase (NAGPA). These mutations account for a small percentage (<10%) of stuttering cases in this study, indicating that still unidentified factors contribute to the disorder. GNPTAB, GNPTG and NAGPA act as enzymes in the lysosomal targeting pathway. Other mutations in GNPTAB and GNPTG are associated with mucolipidoses, disorders associated with deficits in development, mental ability, and speech, though this study is the first to link mutations in these genes to stuttering (Kang et al., 2010; Kang & Drayna, 2012). The mechanisms by which these mutations affect speech are unknown. Other loci have been identified as potential sites for mutations associated with stuttering (Kraft & Yairi, 2012; Raza, Amjad, Riazuddin, & Drayna, 2012), but as yet no other genes have been discovered. One study did find an association between a SNP in the DRD2 gene in a Chinese Han population (Lan et al., 2009), but this result was not replicated in a larger sample (Kang et al., 2011). Additionally, a case was reported in which a partial deletion of CNTNAP2 was found in a stuttering patient, (Petrin et al., 2010) suggesting that there may be some overlap of genetic factors in stuttering and other language disorders.
6. Other Genes of Interest
Additional genes likely contribute to vocal learning. In a screen of genes within a region on chromosome 16 associated with SLI, two candidates correlated with measures of language ability: c-maf-inducing protein (CMIP) and calcium-importing ATPase, type 2C, member 2 (ATP2C2; Newbury et al., 2009). A subsequent study found an association of CMIP, but not of ATP2C2, with reading-related measures (Newbury et al., 2011; Scerri et al., 2011). Though both molecules are expressed in the brain, their functions therein are still poorly understood. In other tissues, CMIP is involved in a cell signaling cascade (Grimbert et al., 2003), and ATP2C2 is part of a pathway responsible for shuttling divalent ions to the Golgi apparatus (Faddy et al., 2008; Missiaen, Dode, Vanoevelen, Raeymaekers, & Wuytack, 2007). Other genes potentially involved in language comprehension include doublecortin domain containing protein 2 (DCDC2) and KIAA0319, which have both been associated with dyslexia (Czamara et al., 2011; Newbury et al., 2011; M. L. Rice, Smith, & Gayán, 2009; Scerri et al., 2011). Recently, DCDC2 was found to affect neuronal firing, increasing the excitability and compromising spike timing (Che, Girgenti, & Loturco, 2013). Given that the other genes implicated in language acquisition and production seem to be involved in either neurogenesis or neurite growth, perhaps CMIP, ATP2C2, and DCDC2 affect either or both of these processes. However, the function of KIAA0319 in language processing is beginning to be better understood. KIAA0319 is involved in the clathrin endocytosis pathway (Levecque, Velayos-Baeza, Holloway, & Monaco, 2009). Knockdown of Kiaa0319 expression in rat auditory cortex results in increased neuronal input resistance accompanied by increased excitability in response to auditory stimuli (Centanni et al., 2013). The authors hypothesize that this change in neuronal excitability, relevant to variants of KIAA0319 in cases of dyslexia, impedes differentiation of speech and non-speech sounds. Another gene of interest in relation to its role in language is FMR1, which encodes the fragile X mental retardation protein (FMRP). Language delay and impairments in both receptive and expressive language are characteristic of children with fragile X syndrome (FXS; Finestack, Richmond, & Abbeduto, 2009). In the zebra finch song system, FMRP is expressed in song nuclei HVC, LMAN, RA, and area X (Winograd, Clayton, & Ceman, 2008). Interestingly, FMRP is enriched in male RA around the onset of the sensorimotor learning phase. These data suggest that FMRP may be a common factor in learned vocalizations in both humans and songbirds.
7. MicroRNA
MicroRNAs (miRs) are short (~22 nucleotide), noncoding RNAs that post-transcriptionally regulate synthesis of specific proteins through either degradation of the mRNA or inhibition of translation (He & Hannon, 2004; Pasquinelli, 2012). These small molecules are thought to “fine-tune” gene expression involved in many biological processes. Research on miR functions in the brain has focused primarily on roles in development and neurogenesis (Liu & Zhao, 2009; Sun, Crabtree, & Yoo, 2013), though studies are starting to emerge on activational effects in the mature brain (Bredy, Lin, Wei, Baker-Andresen, & Mattick, 2011; Fiore, Khudayberdiev, Saba, & Schratt, 2011; Shi et al., 2013). MicroRNAs can affect learning and memory-based tasks, such as fear conditioning, context conditioning, place preference, and Morris water maze performance (Griggs, Young, Rumbaugh, & Miller, 2013; Konopka et al., 2010; Olde Loohuis et al., 2011; Wang & Barres, 2012). Another class of small noncoding RNAs are those that interact with regulatory piwi proteins (piRNAs) in spermatogenic cells, whose mechanisms and functions are still poorly understood, though evidence suggests they are involved in epigenetic control of transcription (Kuramochi-Miyagawa et al., 2008). Recently, piRNAs have been identified as factors contributing to associative learning in Aplysia through regulation of CREB2 (Rajasethupathy et al., 2012). However, investigation into the role of small RNAs in vocal learning has only just begun.
As with many of the genes described in this review, FOXP2 may be used as a starting point by identifying miRs that regulate expression of FOXP2, or are targets of FOXP2 regulation (or in some cases, both). In microarray analysis used to identify gene networks influenced by Foxp2 expression, 22 miRs were identified as transcriptional targets of murine Foxp2 (Vernes et al., 2011). Of these, several have documented functions in the brain: miR-9, -29a, -30a, -30d, -34b, -124a, -125b, and -137. Additional sources of potential vocal learning–associated miRs come from studies in songbirds. In zebra finches, miR-137 was included in a microarray study investigating genes regulated by singing in basal ganglia nucleus area X, and was found to belong to the same gene network module as FoxP2, and negatively associated with the number of motifs sung (Hilliard, Miller, Fraley, et al., 2012). As mentioned in an earlier section, miR-9 and -140-5p are expressed in zebra finch area X, are upregulated by singing in juveniles and adults, and associated with reduced levels of FoxP2 mRNA (Shi et al., 2013). Expression of five miRs in cortical auditory regions are affected by exposure to conspecific song: mir-92, -124, and -129-5p decreased, and mir-25 and -192 increased (Gunaratne et al., 2011). Though the birds in this latter study were adults, and therefore past the critical phase of song learning, the miRs involved in auditory processing may very well impact song learning earlier in life. mir-2954, a putatively avian-specific miR, is expressed at greater levels in males than females in all tissues tested, including brain (Luo et al., 2012). miR-2954 may therefore play a role in the sex-based differences in neuroanatomy and song learning in this species. miRs like miR-2954, which appear to be unique to birds or specifically zebra finch (Gunaratne et al., 2011; Luo et al., 2012), are not likely a common factor underlying behavior in all vocal learning species, although they may regulate genes in a manner common to all vocal learners. A better understanding of the mRNA targets of these miRs will be required to parse out this hypothesis.
How might miRs in the brain affect vocal learning? As with other genes implicated in vocal learning, many miRs act early in development to regulate neurogenesis (Sun et al., 2013), which may contribute to the organization of brain structures underlying speech and vocal learning. In chick spinal cord, miR-9 acts through regulation of FoxP1 to direct motor neuron specification (Otaegi, Pollock, Hong, & Sun, 2011). In the ventricular zone of developing mouse and zebra fish brain, miR-9 promotes neural differentiation by suppression of proteins involved in the proliferation of neural stem cells (Coolen, Thieffry, Drivenes, Becker, & Bally-Cuif, 2012; Saunders et al., 2010; Shibata, Nakao, Kiyonari, Abe, & Aizawa, 2011; Tan, Ohtsuka, González, & Kageyama, 2012; Zhao, Sun, Li, & Shi, 2009). Similarly, miR-124 expression in the developing CNS is thought to direct cell differentiation to a neuronal fate by suppressing non-neuronal transcripts (Cheng, Pastrana, Tavazoie, & Doetsch, 2009; Lim et al., 2005; Makeyev, Zhang, Carrasco, & Maniatis, 2007; Sanuki et al., 2011; Visvanathan, Lee, Lee, Lee, & Lee, 2007). miR-137 also regulates maturation of neurons (Smrt et al., 2010).
Additionally, miRs may have activational effects that support vocal learning. Several miRNAs impact neurite outgrowth and synaptogenesis. miR-9, for example, is expressed in axons of post-mitotic cortical neurons and limits or fine-tunes axon growth (Dajas-Bailador et al., 2012). Brain-derived neurotrophic factor (BDNF) indirectly affects axon growth through regulation of miR-9. Application of BDNF for a short period reduces miR-9 levels and subsequent growth of the axon, but prolonged exposure leads to an increase in miR-9 and a cessation of axon growth. In the songbird, BDNF is thought to be an important factor for neural connectivity between motor song nuclei in development and in adulthood in seasonal learners (Brenowitz, 2013); therefore, miR-9 activity in the songbird brain may be regulated by BDNF exposure. Additionally, predicted binding sites for miR-9 are found in the 3’-untranslated region of matrix metallopeptidase-9 (MMP9), an enzyme that affects synaptic morphology (Konopka et al., 2010). miR-9 represses both Foxp1 (Otaegi et al., 2011) and Foxp2 (Clovis et al., 2012; Shi et al., 2013), whereas Foxp2 promotes miR-9 expression in neuron-like cells in culture (Vernes et al., 2011). This argues for the existence of a Foxp2/miR-9 feedback loop, in which miR-9 indirectly affects gene expression downstream of FoxP2. miR-29a/b changes dendritic spine morphology in hippocampus (Lippi et al., 2011). In Aplysia, miR-124 restricts serotonin-induced synaptic plasticity through regulation of CREB (Rajasethupathy et al., 2009). In mouse differentiating and adult primary cortical neurons, overexpression of miR-124 increases neurite outgrowth, whereas functional blockade causes a delay (Yu, Chung, Deo, Thompson, & Turner, 2008). miRs may affect synaptic plasticity by regulating synaptic molecules. miR-137 has potential binding sites in the 3’UTR of GluR1 mRNA, and miR-124 in GluR2 (Konopka et al., 2010). Regulation of these proteins could impact the synaptic plasticity required for vocal learning.
Conclusions
Recent advances have augmented our understanding of the genetic basis for vocal learning by (a) uncovering new genetic factors through studies of human pathology, (b) discovering new vocal learning–related genes through network analysis of neural tissues pertaining to human speech and birdsong, and (c) developing a better understanding of the physiological effects of known speech-related genes, such as FOXP1, FOXP2, and CNTNAP2 using animal models. FOXP2 was the first gene directly correlated with a language disorder, and through its molecular connections other language-related genes are being discovered, including those in the HGF signaling pathway. As small RNA regulatory factors become better cataloged, we are likely to learn even more about the genetic basis of vocal learning. Since convergent evolution has produced vocal learning in humans, other mammals, and songbirds, we might expect that there are overlapping genes between the clades, but equally we expect some differences. This is likely the case with uPAR, which has no direct avian correlate, but is associated with human speech pathology. Continuing investigation into genes that affect language and vocal learning in other species will provide a better understanding of the mechanisms that govern this complex communicative behavior.
References
Abrahams, B. S., & Geschwind, D. H. (2008). Advances in autism genetics: On the threshold of a new neurobiology. Nature Reviews Genetics, 9(5), 341–355. doi:10.1038/nrg2346
Alarcón, M., Abrahams, B. S., Stone, J. L., Duvall, J. A., Perederiy, J. V., Bomar, J. M., . . . Geschwind, D. H. (2008). Linkage, association, and gene-expression analyses identify CNTNAP2 as an autism-susceptibility gene. American Journal of Human Genetics, 82(1), 150–159. doi:10.1016/j.ajhg.2007.09.005
Ambrose, N. G., Cox, N. J., & Yairi, E. (1997). The genetic basis of persistence and recovery in stuttering. Journal of Speech, Language, and Hearing Research, 40(3), 567–580. doi:10.1044/jslhr.4003.567
Anderson, G. R., Galfin, T., Xu, W., Aoto, J., Malenka, R. C., & Südhof, T. C. (2012). Candidate autism gene screen identifies critical role for cell-adhesion molecule CASPR2 in dendritic arborization and spine development. Proceedings of the National Academy of Sciences. doi:10.1073/pnas.1216398109
Arking, D. E., Cutler, D. J., Brune, C. W., Teslovich, T. M., West, K., Ikeda, M., . . . Chakravarti, A. (2008). A common genetic variant in the neurexin superfamily member CNTNAP2 increases familial risk of autism. American Journal of Human Genetics, 82(1), 160–164. doi:10.1016/j.ajhg.2007.09.015
Arriaga, G., Zhou, E. P., & Jarvis, E. D. (2012). Of mice, birds, and men: The mouse ultrasonic song system has some features similar to humans and song-learning birds. PLoS ONE, 7(10), e46610. doi:10.1371/journal.pone.0046610
Bacon, C., & Rappold, G. A. (2012). The distinct and overlapping phenotypic spectra of FOXP1 and FOXP2 in cognitive disorders. Human Genetics, 131(11), 1687–1698. doi:10.1007/s00439-012-1193-z
Bakkaloglu, B., O’Roak, B. J., Louvi, A., Gupta, A. R., Abelson, J. F., Morgan, T. M., . . . State, M. W. (2008). Molecular cytogenetic analysis and resequencing of contactin associated protein-like 2 in autism spectrum disorders. American Journal of Human Genetics, 82(1), 165–173. doi:10.1016/j.ajhg.2007.09.017
Balmer, T. S., Carels, V. M., Frisch, J. L., & Nick, T. A. (2009). Modulation of perineuronal nets and parvalbumin with developmental song learning. The Journal of Neuroscience, 29(41), 12878–12885. doi:10.1523/JNEUROSCI.2974-09.2009
Benayoun, B. A., Caburet, S., & Veitia, R. A. (2011). Forkhead transcription factors: Key players in health and disease. Trends in Genetics, 27(6), 224–232. doi:10.1016/j.tig.2011.03.003
Benítez-Burraco, A., & Longa, V. M. (2012). Righthandedness, lateralization and language in Neanderthals: A comment on Frayer et al. (2010). Journal of Anthropological Sciences = Rivista di antropologia : JASS / Istituto italiano di antropologia, 90, 187–92– discussion 193–7. doi:10.4436/jass.90002
Berwick, R. C., Friederici, A. D., Chomsky, N., & Bolhuis, J. J. (2013). Evolution, brain, and the nature of language. Trends in Cognitive Sciences, 17(2), 89–98. doi:10.1016/j.tics.2012.12.002
Blasi, F., & Carmeliet, P. (2002). uPAR: A versatile signalling orchestrator. Nature Reviews. Molecular Cell Biology, 3(12), 932–943. doi:10.1038/nrm977
Bottaro, D. P., Rubin, J. S., Faletto, D. L., Chan, A. M., Kmiecik, T. E., Vande Woude, G. F., & Aaronson, S. A. (1991). Identification of the hepatocyte growth factor receptor as the c-met proto-oncogene product. Science (New York, NY), 251(4995), 802–804. doi:10.1126/science.1846706
Bowers, J. M., Perez-Pouchoulen, M., Edwards, N. S., & McCarthy, M. M. (2013). Foxp2 mediates sex differences in ultrasonic vocalization by rat pups and directs order of maternal retrieval. The Journal of Neuroscience, 33(8), 3276–3283. doi:10.1523/JNEUROSCI.0425-12.2013
Bredy, T. W., Lin, Q., Wei, W., Baker-Andresen, D., & Mattick, J. S. (2011). MicroRNA regulation of neural plasticity and memory. Neurobiology of Learning and Memory, 96(1), 89–94. doi:10.1016/j.nlm.2011.04.004
Brenowitz, E. A. (2013). Testosterone and brain-derived neurotrophic factor interactions in the avian song control system. Neuroscience, 239, 115–123. doi:10.1016/j.neuroscience.2012.09.023
Bruce, H. A., & Margolis, R. L. (2002). FOXP2: Novel exons, splice variants, and CAG repeat length stability. Human Genetics, 111(2), 136–144. doi:10.1007/s00439-002-0768-5
Campbell, D. B., D’Oronzio, R., Garbett, K., Ebert, P. J., Mirnics, K., Levitt, P., & Persico, A. M. (2007). Disruption of cerebral cortex MET signaling in autism spectrum disorder. Annals of Neurology, 62(3), 243–250. doi:10.1002/ana.21180
Campbell, D. B., Li, C., Sutcliffe, J. S., Persico, A. M., & Levitt, P. (2008). Genetic evidence implicating multiple genes in the MET receptor tyrosine kinase pathway in autism spectrum disorder. Autism Research: Official Journal of the International Society for Autism Research, 1(3), 159–168. doi:10.1002/aur.27
Campbell, D. B., Sutcliffe, J. S., Ebert, P. J., Militerni, R., Bravaccio, C., Trillo, S., . . . Levitt, P. (2006). A genetic variant that disrupts MET transcription is associated with autism. Proceedings of the National Academy of Sciences of the United States of America, 103(45), 16834–16839. doi:10.1073/pnas.0605296103
Campbell, D. B., Warren, D., Sutcliffe, J. S., Lee, E. B., & Levitt, P. (2010). Association of MET with social and communication phenotypes in individuals with autism spectrum disorder. American Journal of Medical Genetics. Part B, Neuropsychiatric Genetics: The Official Publication of the International Society of Psychiatric Genetics, 153B(2), 438–446. doi:10.1002/ajmg.b.30998
Carlsson, P., & Mahlapuu, M. (2002). Forkhead transcription factors: Key players in development and metabolism. Developmental Biology, 250(1), 1–23. doi:10.1006/dbio.2002.0780
Carr, C. W., Moreno-De-Luca, D., Parker, C., Zimmerman, H. H., Ledbetter, N., Martin, C. L., . . . Abdul-Rahman, O. A. (2010). Chiari I malformation, delayed gross motor skills, severe speech delay, and epileptiform discharges in a child with FOXP1 haploinsufficiency.
European Journal of Human Genetics, 18(11), 1216–1220. doi:10.1038/ejhg.2010.96
Centanni, T. M., Booker, A. B., Sloan, A. M., Chen, F., Maher, B. J., Carraway, R. S., . . . Kilgard, M. P. (2013). Knockdown of the dyslexia-associated gene Kiaa0319 impairs temporal responses to speech stimuli in rat primary auditory cortex. Cerebral Cortex (New York, NY: 1991). doi:10.1093/cercor/bht028
Che, A., Girgenti, M. J., & Loturco, J. (2013). The dyslexia-associated gene Dcdc2 is required for spiketiming precision in mouse neocortex. Biological Psychiatry. doi:10.1016/j.biopsych.2013.08.018
Cheng, L.-C., Pastrana, E., Tavazoie, M., & Doetsch, F. (2009). miR-124 regulates adult neurogenesis in the subventricular zone stem cell niche. Nature Neuroscience, 12(4), 399–408. doi:10.1038/nn.2294
Clovis, Y. M., Enard, W., Marinaro, F., Huttner, W. B., & De Pietri Tonelli, D. (2012). Convergent repression of Foxp2 3’UTR by miR-9 and miR-132 in embryonic mouse neocortex: Implications for radial migration of neurons. Development (Cambridge, England), 139(18), 3332–3342. doi:10.1242/dev.078063
Condro, M. C., & White, S. A. (2014). Distribution of language-related Cntnap2 protein in neural circuits critical for vocal learning. The Journal of Comparative Neurology, (522), 169–185. doi:10.1002/cne.23394
Coolen, M., Thieffry, D., Drivenes, Ø., Becker, T. S., & Bally-Cuif, L. (2012). miR-9 controls the timing of neurogenesis through the direct inhibition of antagonistic factors. Developmental Cell, 22(5), 1052–1064. doi:10.1016/j.devcel.2012.03.003
Czamara, D., Bruder, J., Becker, J., Bartling, J., Hoffmann, P., Ludwig, K. U., . . . Schulte-Körne, G. (2011). Association of a rare variant with mismatch negativity in a region between KIAA0319 and DCDC2 in dyslexia. Behavior Genetics, 41(1), 110–119. doi:10.1007/s10519-010-9413-6
Dajas-Bailador, F., Bonev, B., Garcez, P., Stanley, P., Guillemot, F., & Papalopulu, N. (2012). microRNA-9 regulates axon extension and branching by targeting Map1b in mouse cortical neurons. Nature Neuroscience, 15, 697-699. doi:10.1038/nn.3082
Doupe, A. J., & Kuhl, P. K. (1999). Birdsong and human speech: Common themes and mechanisms. Annual Review of Neuroscience, 22, 567–631. doi:10.1146/annurev.neuro.22.1.567
Eagleson, K. L., Bonnin, A., & Levitt, P. (2005). Region- and age-specific deficits in ?-aminobutyric acidergic neuron development in the telencephalon of theuPAR-/- mouse. The Journal of Comparative Neurology, 489(4), 449–466. doi:10.1002/cne.20647
Eagleson, K. L., Campbell, D. B., Thompson, B. L., Bergman, M. Y., & Levitt, P. (2011). The autism risk genes MET and PLAUR differentially impact cortical development. Autism Research: Official Journal of the International Society for Autism Research, 4(1), 68–83. doi:10.1002/aur.172
Enard, W., Gehre, S., Hammerschmidt, K., Holter, S.M., Blass, T., Somel, M., . . . Pääbo, S. (2009). A humanized version of Foxp2 affects cortico-basal ganglia circuits in mice. Cell, 137(5), 961–971. doi:10.1016/j.cell.2009.03.041
Enard, W., Przeworski, M., Fisher, S. E., Lai, C. S. L., Wiebe, V., Kitano, T., . . . Pääbo, S. (2002). Molecular evolution of FOXP2, a gene involved in speech and language. Nature, 418(6900), 869–872. doi:10.1038/nature01025
Faddy, H. M., Smart, C. E., Xu, R., Lee, G. Y., Kenny, P. A., Feng, M., . . . Monteith, G. R. (2008). Localization of plasma membrane and secretory calcium pumps in the mammary gland. Biochemical and Biophysical Research Communications, 369(3), 977–981. doi:10.1016/j.bbrc.2008.03.003
Falivelli, G., De Jaco, A., Favaloro, F. L., Kim, H., Wilson, J., Dubi, N., . . . Comoletti, D. (2012). Inherited genetic variants in autism-related CNTNAP2 show perturbed trafficking and ATF6 activation. Human Molecular Genetics. doi:10.1093/hmg/dds320
Ferland, R. D., Cherry, T., Preware, P., Morrisey, E. E., & Walsh, C. (2003). Characterization of Foxp 2 and Foxp 1 mRNA and protein in the developing and mature brain. The Journal of Comparative Neurology. doi:10.1002/cne.10654
Feuk, L., Kalervo, A., Lipsanen-Nyman, M., Skaug, J., Nakabayashi, K., Finucane, B., . . . Hannula-Jouppi, K. (2006). Absence of a paternally inherited FOXP2 gene in developmental verbal dyspraxia. American Journal of Human Genetics, 79(5), 965-972. doi:10.1086/508902
Finestack, L. H., Richmond, E. K., & Abbeduto, L. (2009). Language development in individuals with fragile X syndrome. Topics in Language Disorders, 29(2), 133–148. doi:10.1097/TLD.0b013e3181a72016
Fiore, R., Khudayberdiev, S., Saba, R., & Schratt, G. (2011). MicroRNA function in the nervous system. Progress in Molecular Biology and Translational Science, 102, 47–100. doi:10.1016/B978-0-12-415795-8.00004-0
Fisher, S. E., Vargha-Khadem, F., Watkins, K. E., Monaco, A. P., & Pembrey, M. E. (1998). Localisation of a gene implicated in a severe speech and language disorder. Nature Genetics, 18(2), 168–170. doi:10.1038/ng0298-168
Fitch, W. (2000). The evolution of speech: A comparative review. Trends in Cognitive Sciences, 4(7), 258–267. doi:10.1016/S1364-6613(00)01494-7
Fitch, W. T. (2011). The evolution of syntax: An exaptationist perspective. Frontiers in Evolutionary Neuroscience, 3, 9. doi:10.3389/fnevo.2011.00009
Fitch, W. T. (2012). Evolutionary developmental biology and human language evolution: Constraints on adaptation. Evolutionary Biology, 39(4), 613–637. doi:10.1007/s11692-012-9162-y
French, C. A., Jin, X., Campbell, T. G., Gerfen, E., Groszer, M., Fisher, S. E., & Costa, R. M. (2012). An aetiological Foxp2 mutation causes aberrant striatal activity and alters plasticity during skill learning. Molecular Psychiatry, 17(11), 1077–1085.doi:10.1038/mp.2011.105
Fujita, E., Tanabe, Y., Shiota, A., Ueda, M., Suwa, K., Momoi, M. Y., & Momoi, T. (2008). Ultrasonic vocalization impairment of Foxp2 (R552H) knockin mice related to speech-language disorder and abnormality of Purkinje cells. Proceedings of the National Academy of Sciences of the United States of America, 105(8), 3117–3122. doi:10.1073/pnas.0712298105
Gaub, S., Groszer, M., Fisher, S. E., & Ehret, G. (2010). The structure of innate vocalizations in Foxp2-deficient mouse pups. Genes, Brain, and Behavior, 9(4), 390–401. doi:10.1111/j.1601-183X.2010.00570.x
Green, R. E., Krause, J., Briggs, A. W., Maricic, T., Stenzel, U., Kircher, M., . . . Pääbo, S. (2010). A draft sequence of the Neandertal genome. Science (New York, NY), 328(5979), 710–722. doi:10.1126/science.1188021
Griggs, E. M., Young, E. J., Rumbaugh, G., & Miller, C. A. (2013). MicroRNA-182 regulates amygdaladependent memory formation. The Journal of Neuroscience, 33(4), 1734–1740. doi:10.1523/JNEUROSCI.2873-12.2013
Grimbert, P., Valanciute, A., Audard, V., Pawlak, A., Le gouvelo, S., Lang, P., . . . Sahai, D. (2003). Truncation of C-mip (Tc-mip), a new proximal signaling protein, induces c-maf Th2 transcription factor and cytoskeleton reorganization. The Journal of Experimental Medicine, 198(5), 797–807. doi:10.1084/jem.20030566
Groszer, M., Keays, D. A., Deacon, R. M. J., de Bono, J. P., Prasad-Mulcare, S., Gaub, S., . . . Fisher, S. E. (2008). Impaired synaptic plasticity and motor learning in mice with a point mutation implicated in human speech deficits. Current Biology, 18(5), 354–362. doi:10.1016/j.cub.2008.01.060
Gunaratne, P. H., Lin, Y.-C., Benham, A. L., Drnevich, J., Coarfa, C., Tennakoon, J. B., . . . Clayton, D. F. (2011). Song exposure regulates known and novel microRNAs in the zebra finch auditory forebrain. BMC Genomics, 12(1), 277. doi:10.1186/1471-2164-12-277
Haesler, S., Rochefort, C., Georgi, B., Licznerski, P., Osten, P., & Scharff, C. (2007). Incomplete and inaccurate Vocal imitation after knockdown of FoxP2 in songbird basal ganglia nucleus area X. PLoS Biology, 5(12), e321. doi:10.1371/journal. pbio.0050321
Hamdan, F. F., Daoud, H., Rochefort, D., Piton, A., Gauthier, J., Langlois, M., . . . Michaud, J. L. (2010). De novo mutations in FOXP1 in cases with intellectual disability, autism, and language impairment. American Journal of Human Genetics, 87(5), 671–678. doi:10.1016/j.ajhg.2010.09.017
He, L., & Hannon, G. J. (2004). MicroRNAs: Small RNAs with a big role in gene regulation. Nature Reviews Genetics, 5(7), 522–531. doi:10.1038/nrg1379
Hilliard, A. T., Miller, J. E., Fraley, E. R., Horvath, S., & White, S. A. (2012). Molecular microcircuitry underlies functional specification in a Basal Ganglia circuit dedicated to vocal learning. Neuron, 73(3), 537–552. doi:10.1016/j.neuron.2012.01.005
Hilliard, A. T., Miller, J. E., Horvath, S., & White, S. A. (2012). Distinct neurogenomic states in basal ganglia subregions relate differently to singing behavior in songbirds. PLoS Computational Biology, 8(11), e1002773. doi:10.1371/journal.pcbi.1002773
Hockemeyer, D., Sfeir, A. J., Shay, J. W., Wright, W. E., & de Lange, T. (2005). POT1 protects telomeres from a transient DNA damage response and determines how human chromosomes end. The EMBO Journal, 24(14), 2667–2678. doi:10.1038/sj.emboj.7600733
Horn, D. (2012). Mild to moderate intellectual disability and significant speech and language deficits in patients with FOXP1 deletions and mutations. Molecular Syndromology, 2(3–5), 213–216. doi:10.1159/000330916
Horn, D., Kapeller, J., Rivera-Brugués, N., Moog, U., Lorenz-Depiereux, B., Eck, S., . . . Strom, T. M. (2010). Identification of FOXP1 deletions in three unrelated patients with mental retardation and significant speech and language deficits. Human Mutation, 31(11), E1851–60. doi:10.1002/humu.21362
Immelmann, K. (1969). Song development in the zebra finch and other estrildid finches. In R. A. Hinde (Ed.), Bird Vocalizations (pp. 61–74). New York: Cambridge University Press.
Jarvis, E. D. (2004). Learned birdsong and the neurobiology of human language. Annals of the New York Academy of Sciences, 1016(1), 749–777. doi:10.1196/annals.1298.038
Judson, M. C., Bergman, M. Y., Campbell, D. B., Eagleson, K. L., & Levitt, P. (2009). Dynamic gene and protein expression patterns of the autism-associated met receptor tyrosine kinase in the developing mouse forebrain. Journal of Comparative Neurology, 513(5), 511–531. doi:10.1002/cne.21969
Kaestner, K. H., Knochel, W., & Martinez, D. E. (2000). Unified nomenclature for the winged helix/forkhead transcription factors. Genes & Development, 14(2), 142–146. doi:10.1101/gad.14.2.142
Kang, C., Domingues, B. S., Sainz, E., Domingues, C. E. F., Drayna, D., & Moretti-Ferreira, D. (2011). Evaluation of the association between polymorphisms at the DRD2 locus and stuttering. Journal of Human Genetics, 56(6), 472–473. doi:10.1038/jhg.2011.29
Kang, C., & Drayna, D. (2012). A role for inherited metabolic deficits in persistent developmental stuttering. Molecular Genetics and Metabolism, 107(3), 276–280. doi:10.1016/j.ymgme.2012.07.020
Kang, C., Riazuddin, S., Mundorff, J., Krasnewich, D., Friedman, P., Mullikin, J. C., & Drayna, D. (2010). Mutations in the lysosomal enzyme-targeting pathway and persistent stuttering. The New England Journal of Medicine, 362(8), 677–685. doi:10.1056/NEJMoa0902630
Kiya,
T., Itoh, Y., & Kubo, T. (2008). Expression analysis of the FoxP homologue in the brain of the honeybee, Apis mellifera. (1), 53–60. doi:10.1111/j.1365-2583.2008.00775.x
Knornschild, M., Nagy, M., Metz, M., Mayer, F., & von Helversen, O. (2010). Complex vocal imitation during ontogeny in a bat. Biology Letters, 6(2), 156–159. doi:10.1098/rsbl.2009.0685
Kojima, S., Kao, M. H., & Doupe, A. J. (2013). Task-related “cortical” bursting depends critically on basal ganglia input and is linked to vocal plasticity. Proceedings of the National Academy of Sciences, 110(12), 4756–4761. doi:10.1073/pnas.1216308110
Konishi, M., & Akutagawa, E. (1985). Neuronal growth, atrophy and death in a sexually dimorphic song nucleus in the zebra finch brain. Nature, 315(6015), 145–147. doi:10.1038/315145a0
Konopka, G., Wexler, E., Rosen, E., Mukamel, Z., Osborn, G. E., Chen, L., . . . Geschwind, D. H. (2012). Modeling the functional genomics of autism using human neurons. Molecular Psychiatry, 17(2), 202–214. doi:10.1038/mp.2011.60
Konopka, W., Kiryk, A., Novak, M., Herwerth, M., Parkitna, J. R., Wawrzyniak, M., . . . Schütz, G. (2010). MicroRNA loss enhances learning and memory in mice. The Journal of Neuroscience, 30(44), 14835–14842. doi:10.1523/JNEUROSCI.3030-10.2010
Kos, M., van den Brink, D., Snijders, T. M., Rijpkema, M., Franke, B., Fernandez, G., & Hagoort, P. (2012). CNTNAP2 and language processing in healthy individuals as measured with ERPs. PLoS ONE, 7(10), e46995. doi:10.1371/journal.pone.0046995
Kraft, S. J., & Yairi, E. (2012). Genetic bases of stuttering: The state of the art, 2011. Folia phoniatrica et logopaedica: Official Organ of the International Association of Logopedics and Phoniatrics (IALP), 64(1), 34–47. doi:10.1159/000331073
Krause, J., Lalueza-Fox, C., Orlando, L., Enard, W., Green, R. E., Burbano, H. A., . . . Pääbo, S. (2007). The derived FOXP2 variant of modern humans was shared with Neandertals. Current Biology, 17(21), 1908–1912. doi:10.1016/j.cub.2007.10.008
Kuramochi-Miyagawa, S., Watanabe, T., Gotoh, K., Totoki, Y., Toyoda, A., Ikawa, M., . . . Nakano, T. (2008). DNA methylation of retrotransposon genes is regulated by Piwi family members MILI and MIWI2 in murine fetal testes. Genes & Development, 22(7), 908–917. doi:10.1101/gad.1640708
Lai, C. S., Fisher, S. E., Hurst, J. A., Vargha-Khadem, F., & Monaco, A. P. (2001). A forkhead-domain gene is mutated in a severe speech and language disorder. Nature, 413(6855), 519–523. doi:10.1038/35097076
Lan, J., Song, M., Pan, C., Zhuang, G., Wang, Y., Ma, W., . . . Wang, W. (2009). Association between dopaminergic genes (SLC6A3 and DRD2) and stuttering among Han Chinese. Journal of Human Genetics, 54(8), 457–460. doi:10.1038/jhg.2009.60
Lazaro, M. T., Penagarikano, O., Dong, H., Geschwind, D. H., & Golshani, P. (2012, October 15). Excitatory/inhibitory imbalance in the mPFC of the Cntnap2 mouse model of autism. Society for Neuroscience. New Orleans, LA.
Levecque, C., Velayos-Baeza, A., Holloway, Z. G., & Monaco, A. P. (2009). The dyslexia-associated protein KIAA0319 interacts with adaptor protein 2 and follows the classical clathrin-mediated endocytosis pathway. American Journal of Physiology. Cell Physiology, 297(1), C160–168. doi:10.1152/ajpcell.00630.2008
Li, G., Wang, J., Rossiter, S. J., Jones, G., & Zhang, S. (2007). Accelerated FoxP2 evolution in echolocating bats. PLoS ONE, 2(9), e900. doi:10.1371/journal.pone.0000900
Li, S., Weidenfeld, J., & Morrisey, E. E. (2004). Transcriptional and DNA binding activity of the Foxp1/2/4 family is modulated by heterotypic and homotypic protein interactions. Molecular and Cellular Biology, 24(2), 809–822. doi:10.1128/MCB.24.2.809-822.2004
Lim, L. P., Lau, N. C., Garrett-Engele, P., Grimson, A., Schelter, J. M., Castle, J., . . . Johnson, J. M. (2005). Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature, 433(7027), 769–773. doi:10.1038/nature03315
Lippi, G., Steinert, J. R., Marczylo, E. L., D’Oro, S., Fiore,
R., Forsythe, I. D., . . . Young, K. W. (2011). Targeting of the Arpc3 actin nucleation factor by miR-29a/b regulates dendritic spine morphology. The Journal of Cell Biology, 194(6), 889–904. doi:10.1083/jcb.201103006
Liu, C., & Zhao, X. (2009). MicroRNAs in adult and embryonic neurogenesis. NeuroMolecular Medicine, 11(3), 141–152. doi:10.1007/s12017-009-8077-y
Lu, M. M., Li, S., Yang, H., & Morrisey, E. E. (2002). Foxp4: A novel member of the Foxp subfamily of winged-helix genes co-expressed with Foxp1 and Foxp2 in pulmonary and gut tissues. Mechanisms of Development, 119 Suppl 1, S197–202. doi:10.1016/S0925-4773(03)00116-3
Luo, G.-Z., Hafner, M., Shi, Z., Brown, M., Feng, G.-H., Tuschl, T., . . . Li, X. (2012). Genome-wide annotation and analysis of zebra finch microRNA repertoire reveal sex-biased expression. BMC Genomics, 13(1), 727. doi:10.1186/1471-2164-13-727
Lyon, G. R., Shaywitz, S. E., & Shaywitz, B. A. (2003). A definition of dyslexia. Annals of Dyslexia, 53(1), 1–14. doi:10.1007/s11881-003-0001-9
MacDermot, K. D., Bonora, E., Sykes, N., Coupe, A.-M., Lai, C. S. L., Vernes, S. C., . . . Fisher, S. E. (2005). Identification of FOXP2 truncation as a novel cause of developmental speech and language deficits. American Journal of Human Genetics, 76(6), 1074–1080. doi:10.1086/430841
Mahrt, E. J., Perkel, D. J., Tong, L., Rubel, E. W., & Portfors, C. V. (2013). Engineered deafness reveals that mouse courtship vocalizations do not require auditory experience. The Journal of Neuroscience, 33(13), 5573–5583. doi:10.1523/JNEUROSCI.5054-12.2013
Makeyev, E. V., Zhang, J., Carrasco, M. A., & Maniatis, T. (2007). The microRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Molecular Cell, 27(3), 435–448. doi:10.1016/j.molcel.2007.07.015
Mars, W. M., Zarnegar, R., & Michalopoulos, G. K. (1993). Activation of hepatocyte growth factor by the plasminogen activators uPA and tPA. The American Journal of Pathology, 143(3), 949–958.
Marui, T., Koishi, S., Funatogawa, I., Yamamoto, K., Matsumoto, H., Hashimoto, O., . . . Sasaki, T. (2005). No association of FOXP2 and PTPRZ1 on 7q31 with autism from the Japanese population. Neuroscience Research, 53(1), 91–94. doi:10.1016/j.neures.2005.05.003
McRae, P. A., Rocco, M. M., Kelly, G., Brumberg, J. C., & Matthews, R. T. (2007). Sensory deprivation alters aggrecan and perineuronal net expression in the mouse barrel cortex. The Journal of Neuroscience, 27(20), 5405–5413. doi:10.1523/JNEUROSCI.5425-06.2007
Miller, J. E., Spiteri, E., Condro, M. C., Dosumu-Johnson, R. T., Geschwind, D. H., & White, S. A. (2008). Birdsong decreases protein levels of FoxP2, a molecule required for human speech. Journal of Neurophysiology, 100(4), 2015–2025. doi:10.1152/jn.90415.2008
Missiaen, L., Dode, L., Vanoevelen, J., Raeymaekers, L., & Wuytack, F. (2007). Calcium in the Golgi apparatus. Cell Calcium, 41(5), 405–416. doi:10.1016/j.ceca.2006.11.001
Mizutani, A., Matsuzaki, A., Momoi, M. Y., Fujita, E., Tanabe, Y., & Momoi, T. (2007). Intracellular distribution of a speech/language disorder associated FOXP2 mutant. Biochemical and Biophysical Research Communications, 353(4), 869–874. doi:10.1016/j.bbrc.2006.12.130
Mukamel, Z., Konopka, G., Wexler, E., Osborn, G. E., Dong, H., Bergman, M. Y., . . . Geschwind, D. H. (2011). Regulation of MET by FOXP2, genes implicated in higher cognitive dysfunction and autism risk. The Journal of Neuroscience, 31(32), 11437–11442. doi:10.1523/JNEUROSCI.0181-11.2011
Newbury, D. F., Bonora, E., Lamb, J. A., Fisher, S. E., Lai, C. S. L., Baird, G., . . . International Molecular Genetic Study of Autism Consortium. (2002). FOXP2 is not a major susceptibility gene for autism or specific language impairment. American Journal of Human Genetics, 70(5), 1318–1327. doi:10.1086/339931
Newbury, D. F., Paracchini, S., Scerri, T. S., Winchester, L., Addis, L., Richardson, A. J., . . . Monaco, A. P. (2011). Investigation of dyslexia and SLI risk variants in reading- and language-impaired subjects. Behavior Genetics, 41(1), 90–104. doi:10.1007/s10519-010-9424-3
Newbury, D. F., Winchester, L., Addis, L., Paracchini, S., Buckingham, L.-L., Clark, A., . . . Monaco, A. P. (2009). CMIP and ATP2C2 modulate phonological short-term memory in language impairment. American Journal of Human Genetics, 85(2), 264–272. doi:10.1016/j.ajhg.2009.07.004
Nottebohm, F., & Arnold, A. P. (1976). Sexual dimorphism in vocal control areas of the songbird brain. Science, 194(4261), 211–213. doi:10.1126/science.959852
Olde Loohuis, N. F. M., Kos, A., Martens, G. J. M., Bokhoven, H., Nadif Kasri, N., & Aschrafi, A. (2011). MicroRNA networks direct neuronal development and plasticity. Cellular and Molecular Life Sciences, 69(1), 89–102. doi:10.1007/s00018-011-0788-1
O’Roak, B. J., Deriziotis, P., Lee, C., Vives, L., Schwartz, J. J., Girirajan, S., . . . Eichler, E. E. (2011). Exome sequencing in sporadic autism spectrum disorders identifies severe de novo mutations. Nature Genetics, 43(6), 585–589. doi:10.1038/ng.835
Otaegi, G., Pollock, A., Hong, J., & Sun, T. (2011). MicroRNA miR-9 modifies motor neuron columns by a tuning regulation of FoxP1 levels in developing spinal cords. The Journal of Neuroscience, 31(3), 809–818. doi:10.1523/JNEUROSCI.4330-10.2011
Owen, K. A., Qiu, D., Alves, J., Schumacher, A. M., Kilpatrick, L. M., Li, J., . . . Ellis, V. (2010). Pericellular activation of hepatocyte growth factor by the transmembrane serine proteases matriptase and hepsin, but not by the membrane-associated protease uPA. The Biochemical Journal, 426(2), 219–228. doi:10.1042/BJ20091448
Palka, C., Alfonsi, M., Mohn, A., Cerbo, R., Guanciali Franchi, P., Fantasia, D., . . . Palka, G. (2012). Mosaic 7q31 deletion involving FOXP2 gene associated with language impairment. Pediatrics, 129(1), e183–188. doi:10.1542/peds.2010-2094
Palumbo, O., D’Agruma, L., Minenna, A. F., Palumbo, P., Stallone, R., Palladino, T., . . . Carella, M. (2013). 3p14.1 de novo microdeletion involving the FOXP1 gene in an adult patient with autism, severe speech delay and deficit of motor coordination. Gene, 516(1), 107–113. doi:10.1016/j.gene.2012.12.073
Panaitof, S. C. (2012). A songbird animal model for dissecting the genetic bases of autism spectrum disorder. Disease Markers, 33(5), 241–249. doi:10.3233/DMA-2012-0918
Panaitof, S. C., Abrahams, B. S., Dong, H., Geschwind, D. H., & White, S. A. (2010). Language-related Cntnap2 gene is differentially expressed in sexually dimorphic song nuclei essential for vocal learning in songbirds. The Journal of Comparative Neurology, 518(11), 1995–2018. doi:10.1002/cne.22318
Pariani, M. J., Spencer, A., Graham, J. M., & Rimoin, D. L. (2009). A 785kb deletion of 3p14.1p13, including the FOXP1 gene, associated with speech delay, contractures, hypertonia and blepharophimosis. European Journal of Medical Genetics, 52(2–3), 123–127. doi:10.1016/j.ejmg.2009.03.012
Pasquinelli, A. E. (2012). MicroRNAs and their targets: recognition, regulation and an emerging reciprocal relationship. Nature Reviews Genetics, 13(4), 271–282. doi:10.1038/nrg3162
Penagarikano, O., Abrahams, B. S., Herman, E. I., Winden, K. D., Gdalyahu, A., Dong, H., . . . Geschwind, D. H. (2011). Absence of CNTNAP2 leads to epilepsy, neuronal migration abnormalities, and core autism-related deficits. Cell, 147(1), 235–246. doi:10.1016/j.cell.2011.08.040
Peter, B., Raskind, W. H., Matsushita, M., Lisowski, M., Vu, T., Berninger, V. W., . . . Brkanac, Z. (2011). Replication of CNTNAP2 association with nonword repetition and support for FOXP2 association with timed reading and motor activities in a dyslexia family sample. Journal of Neurodevelopmental Disorders, 3(1), 39–49. doi:10.1007/s11689-010-9065-0
Petrin, A. L., Giacheti, C. M., Maximino, L. P., Abramides, D. V. M., Zanchetta, S., Rossi, N. F., . . . Murray, J. C. (2010). Identification of a microdeletion at the 7q33-q35 disrupting the CNTNAP2 gene in a Brazilian stuttering case. American Journal of Medical Genetics Part A, 152A(12), 3164–3172. doi:10.1002/ajmg.a.33749
Poliak, S., Salomon, D., Elhanany, H., Sabanay, H., Kiernan, B., Pevny, L., . . . Peles, E. (2003). Juxtaparanodal clustering of Shaker-like K+ channels in myelinated axons depends on Caspr2 and TAG-1. The Journal of Cell Biology, 162(6), 1149–1160. doi:10.1083/jcb.200305018
Powell, E. M., Campbell, D. B., Stanwood, G. D., Davis, C., Noebels, J. L., & Levitt, P. (2003). Genetic disruption of cortical interneuron development causes region- and GABA cell type-specific deficits, epilepsy, and behavioral dysfunction. The Journal of Neuroscience, 23(2), 622–631.
Raca, G., Baas, B. S., Kirmani, S., Laffin, J. J., Jackson, C. A., Strand, E. A., . . . Shriberg, L. D. (2013). Childhood Apraxia of Speech (CAS) in two patients with 16p11.2 microdeletion syndrome. European Journal of Human Genetics, 21(4), 455–459. doi:10.1038/ejhg.2012.165
Rajasethupathy, P., Antonov, I., Sheridan, R., Frey, S., Sander, C., Tuschl, T., & Kandel, E. R. (2012). A role for neuronal piRNAs in the epigenetic control of memory-related synaptic plasticity. Cell, 149(3), 693–707. doi:10.1016/j.cell.2012.02.057
Rajasethupathy, P., Fiumara, F., Sheridan, R., Betel, D., Puthanveettil, S. V., Russo, J. J., . . . Kandel, E. (2009). Characterization of small RNAs in Aplysia reveals a role for miR-124 in constraining synaptic plasticity through CREB. Neuron, 63(6), 803–817. doi:10.1016/j.neuron.2009.05.029
Raza, M. H., Amjad, R., Riazuddin, S., & Drayna, D. (2012). Studies in a consanguineous family reveal a novel locus for stuttering on chromosome 16q. Human Genetics, 131(2), 311–313. doi:10.1007/s00439-011-1134-2
Reich, D., Green, R. E., Kircher, M., Krause, J., Patterson, N., Durand, E. Y., . . . Hublin, J.-J. (2010). Genetic history of an archaic hominin group from Denisova Cave in Siberia. Nature, 468(7327), 1053–1060. doi:10.1038/nature09710
Reimers-Kipping, S., Hevers, W., Pääbo, S., & Enard, W. (2011). Humanized Foxp2 specifically affects corticobasal ganglia circuits. Neuroscience, 175, 75–84. doi:10.1016/j.neuroscience.2010.11.042
Reiner, A., Perkel, D. J., Bruce, L. L., Butler, A. B., Csillag, A., Kuenzel, W., . . . Jarvis, E. D. (2004). Revised nomenclature for avian telencephalon and some related brainstem nuclei. The Journal of Comparative Neurology, 473(3), 377–414. doi:10.1002/cne.20118
Riaz, N., Steinberg, S., Ahmad, J., Pluzhnikov, A., Riazuddin, S., Cox, N. J., & Drayna, D. (2005). Genomewide significant linkage to stuttering on chromosome 12. American Journal of Human Genetics, 76(4), 647–651. doi:10.1086/429226
Rice, G. M., Raca, G., Jakielski, K. J., Laffin, J. J., Iyama-Kurtycz, C. M., Hartley, S. L., . . . Shriberg, L. D. (2012). Phenotype of FOXP2 haploinsufficiency in a mother and son. American Journal of Medical Genetics Part A, 158A(1), 174–181. doi:10.1002/ajmg.a.34354
Rice, M. L., Smith, S. D., & Gayán, J. (2009). Convergent genetic linkage and associations to language, speech and reading measures in families of probands with Specific Language Impairment. Journal of Neurodevelopmental Disorders, 1(4), 264–282. doi:10.1007/s11689-009-9031-x
Rochefort, C., He, X., Scotto-Lomassese, S., & Scharff, C. (2007). Recruitment of FoxP2-expressing neurons to area X varies during song development. Developmental Neurobiology, 67(6), 809–817. doi:10.1002/dneu.20393
Roll, P., Rudolf, G., Pereira, S., Royer, B., Scheffer, I. E., Massacrier, A., . . . Szepetowski, P. (2006). SRPX2 mutations in disorders of language cortex and cognition. Human Molecular Genetics, 15(7), 1195–1207. doi:10.1093/hmg/ddl035
Roll, P., Vernes, S. C., Bruneau, N., Cillario, J., Ponsole- Lenfant, M., Massacrier, A., . . . Szepetowski, P. (2010). Molecular networks implicated in speech-related disorders: FOXP2 regulates the SRPX2/uPAR complex. Human Molecular Genetics, 19(24), 4848–4860. doi:10.1093/hmg/ddq415
Rousso, D. L., Pearson, C. A., Gaber, Z. B., Miquelajauregui, A., Li, S., Portera-Cailliau, C., . . . Novitch, B. G. (2012). Foxp-mediated suppression of N-cadherin regulates neuroepithelial character and progenitor maintenance in the CNS. Neuron, 74(2), 314–330. doi:10.1016/j.neuron.2012.02.024
Royer, B., Soares, D. C., Barlow, P. N., Bontrop, R. E., Roll, P., Robaglia-Schlupp, A., . . . Szepetowski, P. (2007). Molecular evolution of the human SRPX2 gene that causes brain disorders of the Rolandic and Sylvian speech areas. BMC Genetics, 8, 72. doi:10.1186/1471-2156-8-72
Royer-Zemmour, B., Ponsole-Lenfant, M., Gara, H., Roll, P., Lévêque, C., Massacrier, A., . . . Szepetowski, P. (2008). Epileptic and developmental disorders of the speech cortex: Ligand/receptor interaction of wild-type and mutant SRPX2 with the plasminogen activator receptor uPAR. Human Molecular Genetics, 17(23), 3617–3630. doi:10.1093/hmg/ddn256
Santucci, A. (1995). Effects of scopolamine on spatial working memory in rats pretreated with the serotonergic depletor p-Chloroamphetamine. Neurobiology of Learning and Memory, 63(3), 286–290. doi:10.1006/nlme.1995.1033
Sanuki, R., Onishi, A., Koike, C., Muramatsu, R., Watanabe, S., Muranishi, Y., . . . Furukawa, T. (2011). miR-124a is required for hippocampal axogenesis and retinal cone survival through Lhx2 suppression. Nature Neuroscience, 14(9), 1125–1134. doi:10.1038/nn.2897
Saunders, L. R., Sharma, A. D., Tawney, J., Nakagawa, M., Okita, K., Yamanaka, S., . . . Verdin, E. (2010). miRNAs regulate SIRT1 expression during mouse embryonic stem cell differentiation and in adult mouse tissues. Aging, 2(7), 415–431.
Scerri, T. S., Morris, A. P., Buckingham, L.-L., Newbury, D. F., Miller, L. L., Monaco, A. P., . . . Paracchini, S. (2011). DCDC2, KIAA0319 and CMIP are associated with reading-related traits. Biological Psychiatry, 70(3), 237–245. doi:10.1016/j.biopsych.2011.02.005
Scharff, C., & Petri, J. (2011). Evo-devo, deep homology and FoxP2: Implications for the evolution of speech and language. Philosophical Transactions of the Royal Society of London. Series B, Biological Sciences, 366(1574), 2124–2140. doi:10.1098/rstb.2011.0001
Schulz, S. B., Haesler, S., Scharff, C., & Rochefort, C. (2010). Knockdown of FoxP2 alters spine density in Area X of the zebra finch. Genes, Brain, and Behavior, 9(7), 732–740. doi:10.1111/j.1601-183X.2010.00607.x
Scott-Van Zeeland, A. A., Abrahams, B. S., Alvarez-Retuerto, A. I., Sonnenblick, L. I., Rudie, J. D., Ghahremani, D., . . . Bookheimer, S. Y. (2010). Altered functional connectivity in frontal lobe circuits is associated with variation in the autism risk gene CNTNAP2. Science Translational Medicine, 2(56), 56ra80. doi:10.1126/scitranslmed.3001344
Scott-Van Zeeland, A. A., Dapretto, M., Ghahremani, D. G., Poldrack, R. A., & Bookheimer, S. Y. (2010). Reward processing in autism. Autism Research: Official Journal of the International Society for Autism Research, 3(2), 53–67. doi:10.1002/aur.122
Shi, Z., Luo, G., Fu, L., Fang, Z., Wang, X., & Li, X. (2013). miR-9 and miR-140-5p target FoxP2 and are regulated as a function of the social context of singing behavior in zebra finches. The Journal of Neuroscience: The Official Journal of the Society for Neuroscience, 33(42), 16510–16521. doi:10.1523/JNEUROSCI.0838-13.2013
Shibata, M., Nakao, H., Kiyonari, H., Abe, T., & Aizawa, S. (2011). MicroRNA-9 regulates neurogenesis in mouse telencephalon by targeting multiple transcription factors. The Journal of Neuroscience: The Official Journal of the Society for Neuroscience, 31(9), 3407–3422. doi:10.1523/JNEUROSCI.5085-10.2011
Shriberg, L. D., Ballard, K. J., Tomblin, J. B., Duffy, J. R., Odell, K. H., & Williams, C. A. (2006). Speech, prosody, and voice characteristics of a mother and daughter with a 7;13 translocation affecting FOXP2. Journal of Speech, Language, and Hearing Research, 49(3), 500–525. doi:10.1044/1092-4388(2006/038)
Shu, W., Cho, J. Y., Jiang, Y., Zhang, M., Weisz, D., Elder, G. A., . . . Buxbaum, J. D. (2005). Altered ultrasonic vocalization in mice with a disruption in the Foxp2 gene. Proceedings of the National Academy of Sciences of the United States of America, 102(27), 9643–9648. doi:10.1073/pnas.0503739102
Shu, W., Yang, H., Zhang, L., Lu, M. M., & Morrisey, E. E. (2001). Characterization of a new subfamily of winged-helix/forkhead (Fox) genes that are expressed in the lung and act as transcriptional repressors. The Journal of Biological Chemistry, 276(29), 27488–27497. doi:10.1074/jbc.M100636200
Sia, G. M., Clem, R. L., & Huganir, R. L. (2013). The human language-associated gene SRPX2 regulates synapse formation and vocalization in mice. Science (New York, NY), 342(6161), 987–991. doi:10.1126/science.1245079
Smrt, R. D., Szulwach, K. E., Pfeiffer, R. L., Li, X., Guo, W., Pathania, M., . . . Zhao, X. (2010). MicroRNA miR-137 regulates neuronal maturation by targeting ubiquitin ligase mind bomb-1. Stem Cells (Dayton, Ohio), 28(6), 1060–1070. doi:10.1002/stem.431
Stoeger, A. S., Mietchen, D., Oh, S., de Silva, S., Herbst, C. T., Kwon, S., & Fitch, W. T. (2012). An Asian elephant imitates human speech. Current Biology, 22(22), 2144–2148. doi:10.1016/j.cub.2012.09.022
Strauss, K. A., Puffenberger, E. G., Huentelman, M. J., Gottlieb, S., Dobrin, S. E., Parod, J. M., . . . Morton, D. H. (2006). Recessive symptomatic focal epilepsy and mutant contactin-associated protein-like 2. The New England Journal of Medicine, 354(13), 1370–1377. doi:10.1056/NEJMoa052773
Sun, A. X., Crabtree, G. R., & Yoo, A. S. (2013). MicroRNAs: Regulators of neuronal fate. Current Opinion in Cell Biology, 25(2), 215–221. doi:10.1016/j.ceb.2012.12.007
Talkowski, M. E., Rosenfeld, J. A., Blumenthal, I., Pillalamarri, V., Chiang, C., Heilbut, A., . . . Gusella, J. F. (2012). Sequencing chromosomal abnormalities reveals neurodevelopmental loci that confer risk across diagnostic boundaries. Cell, 149(3), 525–537. doi:10.1016/j.cell.2012.03.028
Tan, S.-L., Ohtsuka, T., González, A., & Kageyama, R. (2012). MicroRNA9 regulates neural stem cell differentiation by controlling Hes1 expression dynamics in the developing brain. Genes to Cells: Devoted to Molecular & Cellular Mechanisms, 17(12), 952–961. doi:10.1111/gtc.12009
Tanabe, Y., Fujita, E., & Momoi, T. (2011). FOXP2 promotes the nuclear translocation of POT1, but FOXP2(R553H), mutation related to speech-language disorder, partially prevents it. Biochemical and Biophysical Research Communications, 410(3), 593–596. doi:10.1016/j.bbrc.2011.06.032
Tanaka, K., Arao, T., Tamura, D., Aomatsu, K., Furuta, K., Matsumoto, K., . . . Nishio, K. (2012). SRPX2 is a novel chondroitin sulfate proteoglycan that is overexpressed in gastrointestinal cancer. PLoS ONE, 7(1), e27922. doi:10.1371/journal.pone.0027922
Teramitsu, I., Kudo, L. C., London, S. E., Geschwind, D. H., & White, S. A. (2004). Parallel FoxP1 and FoxP2 expression in songbird and human brain predicts functional interaction. The Journal of Neuroscience, 24(13), 3152–3163. doi:10.1523/JNEUROSCI.5589-03.2004
Teramitsu, I., Poopatanapong, A., Torrisi, S., & White, S. A. (2010). Striatal FoxP2 is actively regulated during songbird sensorimotor learning. PLoS ONE, 5(1), e8548. doi:10.1371/journal.pone.0008548
Teramitsu, I., & White, S. A. (2006). FoxP2 regulation during undirected singing in adult songbirds. The Journal of Neuroscience, 26(28), 7390–7394. doi:10.1523/JNEUROSCI.1662-06.2006
Thompson, C. K., Schwabe, F., Schoof, A., Mendoza, E., Gampe, J., Rochefort, C., & Scharff, C. (2013). Young and intense: FoxP2 immunoreactivity in Area X varies with age, song stereotypy, and singing in male zebra finches. Frontiers in Neural Circuits, 7, 24. doi:10.3389/fncir.2013.00024
Toma, C., Hervás, A., Torrico, B., Balmaña, N., Salgado, M., Maristany, M., . . . Cormand, B. (2013). Analysis of two language-related genes in autism: A casecontrol association study of FOXP2 and CNTNAP2. Psychiatric Genetics, 23(2), 82–85. doi:10.1097/YPG.0b013e32835d6fc6
Tsui, D., Vessey, J. P., Tomita, H., Kaplan, D. R., & Miller, F. D. (2013). FoxP2 regulates neurogenesis during embryonic cortical development. The Journal of Neuroscience, 33(1), 244–258. doi:10.1523/JNEUROSCI.1665-12.2013
Vargha-Khadem, F., Watkins, K., Alcock, K., Fletcher, P., & Passingham, R. (1995). Praxic and nonverbal cognitive deficits in a large family with a genetically transmitted speech and language disorder. Proceedings of the National Academy of Sciences of the United States of America, 92(3), 930–933. doi:10.1073/pnas.92.3.930
Vernes, S. C., MacDermot, K. D., Monaco, A. P., & Fisher, S. E. (2009). Assessing the impact of FOXP1 mutations on developmental verbal dyspraxia. European Journal of Human Genetics, 17(10), 1354–1358. doi:10.1038/ejhg.2009.43
Vernes, S. C., Newbury, D. F., Abrahams, B. S., Winchester, L., Nicod, J., Groszer, M., . . . Phil, D. (2008). A functional genetic link between distinct developmental language disorders. The New England Journal of Medicine, 359(22), 2337–2345. doi:10.1056/NEJMoa0802828
Vernes, S. C., Nicod, J., Elahi, F. M., Coventry, J. A., Kenny, N., Coupe, A.-M., . . . Fisher, S. E. (2006). Functional genetic analysis of mutations implicated in a human speech and language disorder. Human Molecular Genetics, 15(21), 3154–3167. doi:10.1093/hmg/ddl392
Vernes, S. C., Oliver, P. L., Spiteri, E., Lockstone, H. E., Puliyadi, R., Taylor, J. M., . . . Fisher, S. E. (2011). Foxp2 regulates gene networks implicated in neurite outgrowth in the developing brain. PLoS Genetics, 7(7), e1002145. doi:10.1371/journal.pgen.1002145
Visvanathan, J., Lee, S., Lee, B., Lee, J. W., & Lee, S. K. (2007). The microRNA miR-124 antagonizes the antineural REST/SCP1 pathway during embryonic CNS development. Genes & Development, 21(7), 744–749. doi:10.1101/gad.1519107
Wang, B., Lin, D., Li, C., & Tucker, P. (2003). Multiple domains define the expression and regulatory properties of Foxp1 forkhead transcriptional repressors. The Journal of Biological Chemistry, 278(27), 24259–24268. doi:10.1074/jbc.M207174200
Wang, J. T., & Barres, B. A. (2012). Axon degeneration:
Where the Wlds things are. Current Biology, 22(7), R221–3. doi:10.1016/j.cub.2012.02.056
Whalley, H. C., O’Connell, G., Sussmann, J. E., Peel, A., Stanfield, A. C., Hayiou-Thomas, M. E., . . . Hall, J. (2011). Genetic variation in CNTNAP2 alters brain function during linguistic processing in healthy individuals. American Journal of Medical Genetics. Part B, Neuropsychiatric Genetics: The Official Publication of the International Society of Psychiatric Genetics, 156(8), 941–948. doi:10.1002/ajmg.b.31241
Winograd, C., Clayton, D., & Ceman, S. (2008). Expression of fragile X mental retardation protein within the vocal control system of developing and adult male zebra finches. Neuroscience, 157(1), 132–142. doi:10.1016/j.neuroscience.2008.09.005
Yu, J.-Y., Chung, K.-H., Deo, M., Thompson, R. C., & Turner, D. L. (2008). MicroRNA miR-124 regulates neurite outgrowth during neuronal differentiation. Experimental Cell Research, 314(14), 2618–2633. doi:10.1016/j.yexcr.2008.06.002
Zeesman, S., Nowaczyk, M. J. M., Teshima, I., Roberts, W., Cardy, J. O., Brian, J., . . . Scherer, S. W. (2006). Speech and language impairment and oromotor dyspraxia due to deletion of 7q31 that involves FOXP2. American Journal of Medical Genetics Part A, 140(5), 509–514. doi:10.1002/ajmg.a.31110
Zhao, C., Sun, G., Li, S., & Shi, Y. (2009). A feedback regulatory loop involving microRNA-9 and nuclear receptor TLX in neural stem cell fate determination. Nature Structural & Molecular Biology, 16(4), 365–371. doi:10.1038/nsmb.1576
